Chapter: Biotechnology: Recombinant DNA Technology
DNA Library
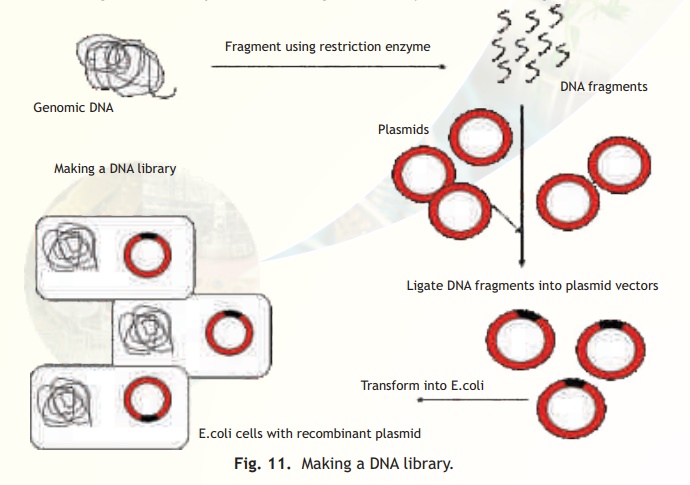
DNA Library
In order to obtain a DNA fragment carrying a useful target gene it is essential to make a DNA library containing all possible DNA fragments from the genome of a given species or cell type. The target gene can then be easily identified from the library and then used for cloning. Remember per cell only two copies (diploid) of a target gene are present and therefore too little is there for detection and use. A DNA library not only contains all possible fragments of DNA from a given cell or organism but also large amounts of the same as a resource. Two types of DNA libraries can be constructed - a genomic DNA library and a cDNA library. A genomic DNA library has all possible DNA sequences in large amounts from the given cell type. A cDNA library on the other hand has only expressed gene sequences such as protein encoding genes. It is obvious that a genomic DNA library is larger than a cDNA library. Whether the fragments are genomic or cDNA (protein encoding) these are inserted into vectors such as plasmids and then introduced into E. colihosts. The cells are diluted and plated on agarose plates so that distinct colonies are formed starting from single cells. Each colony would therefore have an amplified amount of a given fragment. However unlike an E.coli regular library which has a classification system, a DNA library has to be screened for finding a fragment or gene of interest which is a tedious and time taking process. A schematic diagram of the steps in constructing a DNA library is indicated in Fig. 11.
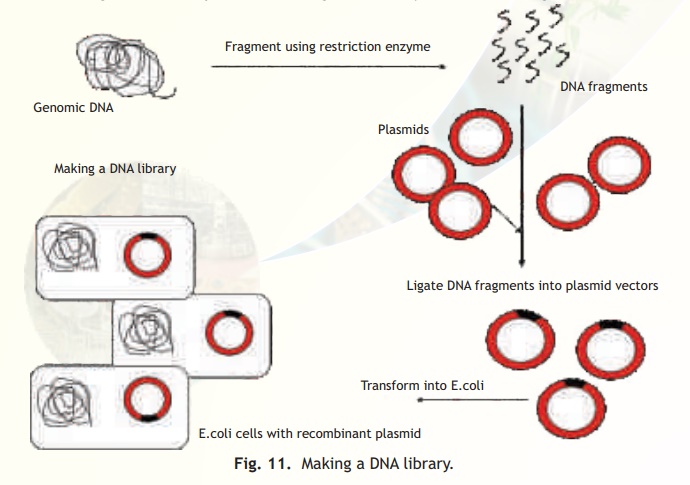
Fig. 11. Making a DNA library.
As shown in Fig. 11, to prepare a genomic library, the total genomic DNA is isolated from a tissue or organism and then fragmented using a restriction enzyme such as EcoRI. An appropriate vector such as pBR322 (plasmid based) is also digested with the same enzyme and is then treated with the enzyme alkaline phosphatase which as indicated elsewhere removes the 5'phosphate group to prevent the plasmid from self ligation. The DNA fragments and cut vector are mixed and then treated with the enzyme DNA ligase. Each vector molecule will contain a different fragment of DNA and these are introduced into E. coli host cells by a technique called transformation. More details about this technique will be discussed in the following section.
Although a genomic library represents the entire genome of an organism, it may not be useful in case of eukaryotic organisms. Genomic DNA of eucaryotes contains non-coding DNA like introns, control regions and repetitive sequences unrecognised by prokaryotic systems such as in E. coli. Therefore another library called cDNA library is preferred especially if the aim of the cloning experiment is to express eukaryotic proteins in an E. coli host. This library has two major advantages over a genomic library. Firstly, it represents only those genes that are being expressed by a particular cell or under specific conditions. Eucaryotes, which include multicellular organisms such as animals and plants have differentiated cells which means that different cells e.g. liver and brain cells express some common proteins and also other proteins which are not common. Secondly, since the source material in constructing such libraries is mRNA, these molecules lack introns and hence would represent only the coding sequences of the genome. However mRNA molecules are highly unstable as they are easily degraded by RNAses and hence these molecules are faithfully copied into the more stable DNA (now called cDNA) before cloning. The construction of a cDNA library begins with the isolation of mRNA from a given cell type or tissue which are copied into cDNA using a special enzyme called reverse transcriptase. The procedure results in double-stranded cDNA which can be incorporated into vectors such as pBR322.
Related Topics