Chapter: Genetics and Molecular Biology: DNA Synthesis
In vitro DNA Replication - Enzymology
In vitro DNA Replication
The mere incorporation of radiolabeled nucleotides
into polymers of DNA is far from the complete biological process of DNA
replication. The initial experiments seeking polymerization activities from
cell extracts used nicked and gapped DNA as a template. This yielded
polymerases capable of elongating DNA, but did not provide an assay for any of
the DNA initiating components. To seek the cellular machinery necessary for
initiating replication, DNA templates were required that contained sequences
specifying origins of replication. The most convenient source of such origins
was small DNA phages since each molecule must contain an origin. The results of
experiments with several different phage templates revealed the astounding fact
that the proteins required for initiating replication varied from one DNA origin
to another. At first it was not possible to discern the biochemical principles
underlying initia-tion of replication. Therefore, when DNA cloning became
possible, attention turned to a replication origin of greater generality and
impor-tance, the origin of replication of the E. coli chromosome.
Later, when it became possible to work with animal viruses and to isolate and
study replication origins from eukaryotic cells, these also were studied.
Kornberg and his collaborators were able to find
conditions in which a cell extract prepared from E. coli could replicate
DNA from the E. coli origin, oriC. Such
an extract undoubtedly possessed many different proteins acting in concert to
replicate the DNA. Once this step was working, it was then possible to seek to
identify specific proteins involved in the reaction. Geneticists assisted this
difficult step through their isolation of temperature-sensitive mutations that
blocked DNA synthesis in growing cells. For example, extracts prepared from
cells with a temperature-sensitive dnaA
mutation were inactive. This, of course, is a biochemist’s dream, for it
provides a specific assay for the DnaA protein. Extracts prepared from
wild-type cells can supplement extracts prepared from temperature-sensitive dnaA mutant cells. This supplementation
results from the wild-type DnaA protein in the wild-type extract. Next, the
wild-type extract can be fractionated and the invitro complementation assay detects which fraction contains the
DnaAprotein. With such an assay the DnaA protein was purified.
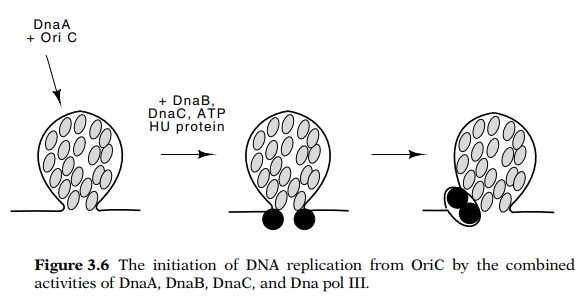
Figure
3.6 The initiation of DNA replication
from OriC by the combinedactivities of DnaA, DnaB, DnaC, and Dna pol III.
The strategy used with the DnaA protein is
straightforward, and it can be used to purify proteins by making use of any
replication mutant whose cell extracts are inactive. The work required for this
approach is immense, and therefore it helps to try to guess proteins required
for replication and to add these as purified components. If the assay doesn’t
replicate DNA, cell extracts are added and the resulting activity can be used
to guide purification of the remaining components. Ultimately, the following
components were identified as necessary for in
vitro replica-tion from oriC:
DnaA protein, DnaB protein, DnaC protein, DnaG protein, DNA polymerase III
holoenzyme, DNA gyrase, single-stranded binding protein, and ATP. Analogous
experiments using replication origins from animal viruses have also permitted
the detection, purifica-tion and study of the complete set of proteins
necessary for their activity.
In vitro initiation and synthesis reactions have permitted
the replica-tion process at the E. coli oriC to be dissected into several
steps. Initiation begins with 20 to 40 molecules of DnaA protein binding to
four sites in the 260 base pair oriC
region (Fig. 3.6). The complex of DnaA and oriC
contains the DNA wrapped around the outside and the protein in the middle. The
large number of protein molecules required
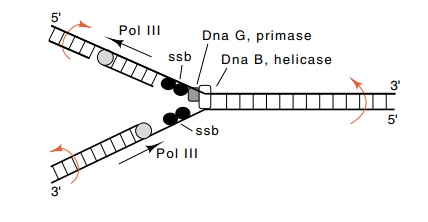
Figure
3.7 The activities in the vicinity of
a DNA replication fork. Also shownare the rotations generated by movement of a
replication fork.
for the first step of initiation makes the step
critically dependent on the concentration of the protein. Such a step is ideal
for tight regulation of the initiation process. In the second step, DnaB plus
DnaC bind. DnaB possesses a helicase activity which separates the DNA strands
with the consumption of energy. DnaA binds to three additional sites in oriC and several regions of
single-stranded DNA are generated. Finally, DnaG protein lays down RNA primers
that are used by the pol III holoenzyme.
During the elongation process the single-stranded
binding protein SSB binds to single-stranded regions opened by the helicase
activities of DnaB (Fig. 3.7). DNA gyrase also is required for elongation. It
untwists the rotations that are generated ahead of the moving replica-tion
fork. In vivo the twists that are
generated behind the moving replication fork are removed by topisomerase I.
Several other proteins are also involved with
replication. The histone-like protein, HU, seems to assist the process, but its
mechanism is unknown. RNAse H digests the RNA from RNA-DNA hybrids left over
from transcription and prevents initiation from points other than the origin.
Related Topics