Chapter: Genetics and Molecular Biology: DNA Synthesis
Gel Electrophoresis Assay of Eukaryotic Replication Origins - Physiological Aspects
Gel Electrophoresis Assay of Eukaryotic
Replication Origins
It has been possible to construct artificial yeast
chromosomes by com-bining isolated centromeres, telomeres, and autonomously
replicating sequences known as ARS sequences. Although ARS sequences function
in the artificial chromosomes, it is interesting to know whether they also
normally function as origins. A gel electrophoresis technique based on the
Southern transfer technology as described permits examination of this question.
Experiments have shown that ordinarily gel electrophoresis separates DNA according to molecular weight. If, however, the voltage gradient is increased five-fold above normal to about 5 V/cm, and higher than normal concentrations of agarose are used, then the separation becomes largely based on shape of the DNA molecules rather than their total molecular weight. Normal electrophoresis and this shape-sensitive electrophoresis can be combined in a two dimensional electrophoretic separation technique that is particularly useful in the analysis of replication origins. The DNA fragments necessary for the analysis are generated by cleaving DNA extracted from cells with restriction enzymes. These cut at specific sequences, and will be discussed. A DNA sample obtained by cutting chromosomal DNA with such a restriction enzyme is first separated according to size by electrophoresis in one direction. Then it is separated according to shape by electrophoresis in a direction perpendicular to the first. After the electrophoresis, the locations of the fragments containing the sequence in question are determined by Southern transfer.
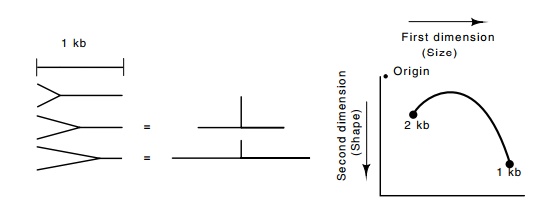
Figure 3.19 The various structures resulting from a replication fork enteringthe 1 kb region from the left. The arc shown indicates the positions of the various structures after 2D electrophoresis.
Let us first consider the two-dimensional electrophoretic pattern of DNA fragments that would be generated from DNA extracted from a large number of growing cells if replication origins were to enter a 1000 base pair region from the left (Fig. 3.19). In the majority of the cells no replication origin will be on the 1000 base pair stretch of DNA. Therefore, after cutting with the restriction enzyme these stretches will be simple 1000 base pair pieces of DNA. After the Southern transfer, these molecules will show up as a spot at 1000 base pairs. Some of the cells would contain replication origins in the 1000 base pair region. After cutting with the restriction enzyme, these molecules not only would possess a mass larger than the 1000 base pair molecules, but also they would be more asymmetric. The extreme of asymmetry would occur in those molecules in which the replication origin was 500 base pairs from the end. These would generate the peak shown in Fig. 3.19. Those molecules for which the replication origin was nearly at the right end would, again, be like simple DNA molecules, but molecules 2000 base pairs long. Thus, the collection of molecular species would generate the arc shown upon two dimensional electrophoresis.
Figure 3.20 The various structures resulting from a replication fork originat-ing from the center of the 1 kb ARS region. The arc of the positions reached by the various structures does not reach the 2 kb point because none of the structures approaches a linear 2 kb molecule.
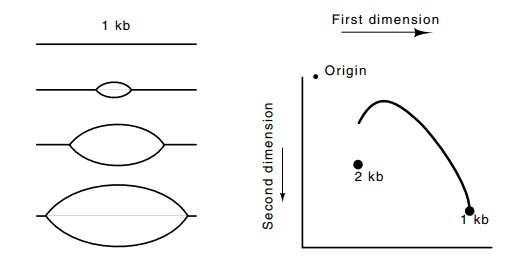
If the region of DNA we are considering contains a
replication origin, quite a different pattern is generated. Suppose the origin
is exactly in the middle of the region. Then the various replication forms are
as shown in Fig. 3.20 and the pattern that will be found after Southern
transfer is as shown. It is left as a problem to deduce the pattern expected if
the origin is located to one side of the center of the DNA segment.
Related Topics