Chapter: Genetics and Molecular Biology: DNA Synthesis
Detection and Basic Properties of DNA Polymerases - Enzymology
Detection and Basic Properties of DNA
Polymerases
Use of a purified enzyme is one requirement for
careful study of DNA synthesis. In the early days of molecular biology,
important experiments by Kornberg demonstrated the existence in cell extracts
of an enzyme that could incorporate nucleoside triphosphates into a DNA chain.
This enzymatic activity could be purified from bacterial extracts, and the
resultant enzyme was available for biochemical studies. Naturally, the first
question to be asked with such an enzyme was whether it utilized a
complementary DNA strand to direct the incorporation of the nucleo-tides into
the elongating strand via Watson-Crick base-pairing rules. Fortunately the
answer was yes. As this enzyme, DNA pol I, was more carefully studied, however,
some of its properties appeared to make it unlikely that the enzyme synthesized
the majority of the cellular DNA.
Cairns tried to demonstrate that pol I was not the key replication enzyme by isolating a bacterial mutant lacking the enzyme. Of course, if the mutant could not survive, his efforts would have yielded nothing. Remarkably, however, he found a mutant with much less than normal activity. Such a result appeared to prove that cells must possess other DNA-synthesizing enzymes, but until a mutant could be found com-pletely lacking DNA pol I, the demonstration was not complete.
Another way to show the existence of the DNA
polymerases other than DNA pol I is to detect and purify them biochemically.
Previously such attempts had proved futile because DNA pol I masked the presence
of other polymerases. Once Cairns’s mutant was available, however, it was a
matter of straightforward biochemistry to examine bacterial extracts for the
presence of additional DNA-polymerizing enzymes. Two more such enzymes were
found: DNA pol II and DNA pol III.
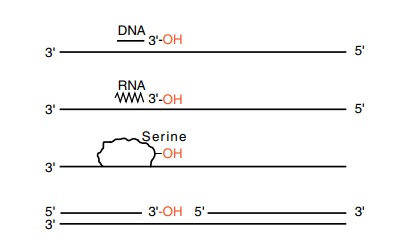
Figure 3.4 Four ways of generating 3’-OH groups necessary for DNA elonga-tion, a deoxyribonucleotide oligomer, a ribonucleotide oligomer, a hydroxyl provided by a protein, and a nick.
None of the three polymerases is capable of
initiating DNA synthesis on a template strand. They are able to elongate
growing polynucleotide chains, but they cannot initiate synthesis of a chain.
This inability is not surprising, however, because initiation must be carefully
regulated and could be expected to involve a number of other proteins that
would not be necessary for elongation. For initiation, all three polymerases
require the presence of a hydroxyl group in the correct position. The hydroxyl
group can derive from a short stretch of DNA or RNA annealed to one strand,
from cleavage of a DNA duplex, or even from a protein, in which the hydroxyl
utilized is on a serine or threonine residue (Fig. 3.4).
In bacteria, DNA pol III is the main DNA
replication enzyme. DNA pol I fills in the gaps in lagging strand synthesis and
assists during repair of damaged DNA. No function is known for DNA pol II. Both
DNA pol I and DNA pol III possess the 3’-to-5’ exonuclease activity that is
necessary for proofreading removal of misincorporated nucleotides.
DNA pol I also possesses a 5’-to-3’ exonuclease
activity not found in the other polymerases. This activity permits pol I to
bind to a nick in the DNA, remove the nucleotide on the 3’ side, incorporate a
nucleotide on the 5’ side and perform this process over and over again without
dissociating from the DNA after each nucleotide. Thus pol I can proc-essively
translate a nick down the DNA in the 5’-to-3’ direction. This process is used
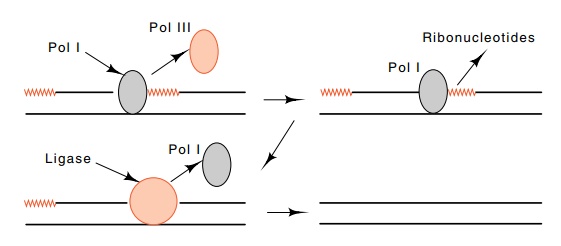
to remove the RNA that primes synthesis of the Okazaki fragments (Fig. 3.5).
When pol III reaches such a primer it dissociates from the DNA. Then DNA pol I
can bind and nick translate through the stretch of RNA. DNA pol I is less
processive than DNA pol III, so that when pol I dissociates at some point after
nick translating through the RNA primer, the resulting DNA-DNA nick can be
sealed by DNA ligase. This ends the elongation process for this Okazaki
fragment.
The two exonuclease activities and the polymerizing
activity of DNA pol I reside on the single polypeptide chain of the enzyme. DNA
pol III is not so simple. Two slightly different forms of the enzyme appear to
function as a dimer, one which synthesizes the leading strand, and one which
synthesizes the lagging strand. Most likely the complete complex, the
holoenzyme, actually consists of at least nine different polypeptide chains.
The α subunit possesses the polymerizing activity, and
the ε subunit contains the 3’-5’ exonuclease activity.
The unit of α, ε, and θ forms an active core that is
capable of synthesizing short stretches of DNA. Remaining associated with the
template strand and being highly processive is conferred by the complex of γ, δ, δ’, χ, and ψ which facilitate the addition of a dimer of β. The dimer of β encircles the DNA and forces the complex to remain associated with the
same DNA molecule through-out the replication cycle.
Not surprisingly, eukaryotic cells also possess
multiple DNA polym - erases which contain multiple subunits. These polymerases,
not the subunits, often are named α, β, γ , δ, and ε. The α
polymerase was long thought to be the general polymerizing enzyme, with the
small β enzyme considered to be a repair enzyme. The γ enzyme is mitochondrial. We now know that α is likely to initiate synthesis on both the
leading and lagging strands since only it possesses priming activity. Further
elonga-tion on the leading strand is performed by δ, and on the lagging strand by ε or α. The β enzyme
could complete the lagging strands by filling the same role as is played by DNA
pol I in bacteria.
Related Topics