Chapter: Medical Microbiology: An Introduction to Infectious Diseases: Viral Multiplication
Genome Replication: DNA, RNA Viruses
GENOME REPLICATION
DNA Viruses
Cells obviously contain the enzymes and accessory proteins required for the replication of DNA. In bacteria these proteins are present continuously, whereas in the eukaryotic cell they are present only during the S phase of the cell cycle, and they are restricted to the nucleus. The extent to which viruses use the cell replication machinery depends on their protein-coding potential and thus on the size of their genome.
The smallest of the DNA viruses, the parvoviruses, are so completely dependent on host machinery that they require the infected cells to be dividing so that a normal S phase will occur and replicate the viral DNA along with the cellular DNA. At the other end of the spectrum are the large DNA viruses, which are relatively independent of cellular func-tions. The largest bacteriophages such as T4 degrade the host cell chromosome early in infection and replace all of the host replication machinery with virus-specified proteins. The largest animal viruses, the poxviruses, are similarly independent of the host. Because they replicate in the cytoplasm, they must code for virtually all of the enzymes and other proteins required for replicating their DNA.
The remainder of the DNA viruses are only partially dependent on host machinery. For example, bacteriophages ΦX174 and code for proteins that direct the initiation of DNA synthesis to the viral origin. However, the actual synthesis of DNA occurs by the complex of cellular enzymes responsible for replication of the Escherichia coli DNA. Similarly the small DNA animal viruses, such as the papovaviruses, code for a protein that is involved in the initiation of synthesis at the origin, but the remainder of the replica-tion process is carried out by host machinery. The somewhat more complex adenoviruses and herpesviruses, in addition to providing origin-specific proteins, also code for their own DNA polymerases and other accessory proteins required for DNA replication.
The fact that the herpesviruses code for their own DNA polymerase has important implications for the treatment of infections by these viruses and illustrates a central principle of antiviral chemotherapy. Certain antiviral drugs (adenine arabinoside and 5 -iododeoxyuridine) have been found to be effective against herpesvirus infections ; they are sufficiently similar to natural substrates that the virally encoded DNA polymerase mistakenly incorporates them into viral DNA, resulting in an inhibition of subsequent DNA synthesis. The host cell enzyme is more discriminating and fails to use the analogs in the synthesis of cellular DNA; thus, the drugs do not kill uninfected cells. The same principle applies to the chain-terminating drugs such as zidovudine (AZT) and dideoxyinosine (ddI) that target the HIV-1 reverse transcriptase. Similarly, the antiviral drugs acyclovir (acycloguanosine) and ganciclovir preferentially kill her-pesvirus-infected cells because the viral nucleoside kinases, unlike the cellular counter-parts, phosphorylate the nucleoside analog, converting it to a form that inhibits further DNA synthesis when DNA polymerases incorporate it into DNA. In principle, any viral process that is distinct from a normal cellular process is a potential target for antiviral drugs. As more becomes known about the details of viral replication, more drugs will be-come available that are targeted to these unique viral processes.
As noted earlier, with the exception of the poxviruses, all of the DNA animal viruses are at least partially dependent on host cell machinery for the replication of their genomes. However, unlike the parvoviruses, the other DNA viruses do not need to infect dividing cells for a productive infection to ensue. Instead, all of these viruses code for a protein expressed early in infection that induces an unscheduled cycle of cellular DNA replication (S phase). In this way, these viruses ensure that the infected cell makes all of the machinery required for the replication of their own DNA. It is noteworthy that all of the DNA viruses except the parvoviruses are capable, in some circumstances, of trans-forming a normal cell into a cancer cell . This correlation suggests that the unlimited proliferative capacity of the cancer cells may be due to the continual synthesis of the viral protein(s) responsible for inducing the unscheduled S phase in a normal infec-tion. The fact that these DNA viruses can induce oncogenic transformation of cell types that are nonpermissive for viral multiplication may simply be an accident related to the need to induce cellular enzymes required for DNA replication during the lytic infection.
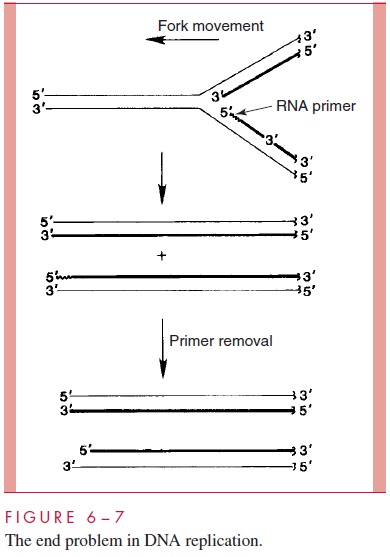
All DNA polymerases, including those encoded by viruses, synthesize DNA chains by the successive addition of nucleotides onto the 3’ end of the new DNA strand. More-over, all DNA polymerases require a primer terminus containing a free 3 -hydroxyl to ini-tiate the synthesis of a DNA chain. In cellular replication, a temporary primer is provided in the form of a short RNA molecule. This primer RNA is synthesized by an RNA poly-merase, and after elongation by the DNA polymerase it is removed. With circular chro-mosomes, such as those found in bacteria and many viruses, the unidirectional chain growth and primer requirement of the DNA polymerase pose no structural problems for replication. However, as illustrated in Figure 6 – 7, when a replication fork encounters the end of a linear DNA molecule, one of the new chains (heavy lines) cannot be completed at its 5’ end, because there exists no means of starting the DNA portion of the chain ex-actly at the end of the template DNA. Thus, after the RNA primer is removed, the new chain is incomplete at its 5’ end. This constraint on the completion of DNA chains on a linear template is called the end problem in DNA replication. Some eukaryotic cells add short repetitive sequences to chromosome ends using an enzyme called telomerase to pre-vent the shortening of the DNA with each successive round of replication.
Several viruses are faced with the end problem during replication of their linear genomes, but none use the cellular telomerase to synthesize DNA ends. It is beyond the scope of this text to detail all of the strategies viruses have evolved to deal with the end problem, but it is worth mentioning some of the structural features found in linear viral genomes whose presence is related to solutions of the end problem. These structures are diagrammed schematically in Figure 6 – 8.
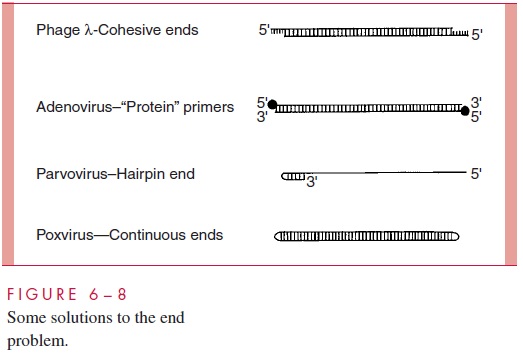
The linear double-stranded genome of bacteriophage possesses 12-bp single-stranded extensions that are complementary in sequence to each other and thus called cohesive ends. Very early after entry into the cell, the two ends pair up to convert the linear genome into a circular molecule to avoid the end problem in replication. The linear double-stranded adenovirus genome contains a protein molecule covalently attached to the 5’ end of both strands. These proteins provide the primers required to initiate the synthesis of the DNA chains during replication, cir-cumventing the need for RNA primers and thus solving the end problem in replication. The single-stranded parvovirus genome contains a self-complementary sequence at the 3’ end that causes the molecule to fold into a hairpin, making it self-priming for DNA replication. The poxviruses contain linear double-stranded genomes in which the ends are continuous. With the parvovirus and poxvirus genomes, the solutions to the end problem create additional problems that must be solved to produce replication products that are identical to the starting genomes.
RNA Viruses
Because nuclear functions are primarily designed for DNA metabolism, RNA animal viruses generally replicate in the cytoplasm. Moreover, cells do not have RNA poly-merases that can copy RNA templates. Therefore, RNA viruses not only need to code for transcriptases, as discussed earlier, but also must provide the replicases required to duplicate the genome RNA.
Furthermore, except in the cases of the RNA phage and the picornaviruses, where transcription and replication are synonymous, the RNA viruses must temporally and functionally separate replication from transcription. This require-ment is especially apparent for the rhabdoviruses, paramyxoviruses, togaviruses, and coronaviruses, where a complete genome, or complementary copy of the genome, is tran-scribed into a set of small monocistronic mRNAs early in infection. After replication be-gins, these same templates are used to synthesize full-length strands for replication.
Two mechanisms exist to separate the process of replication from transcription. First, in some cases, transcription is restricted to subviral particles and involves a transcriptase trans-ported into the cell within the virion. Second, in other cases, the replication process involves either a functionally distinct RNA polymerase or depends on the presence of some other viral-specific accessory protein that directs the synthesis of full-length copies of the tem-plate rather than the shorter monocistronic mRNAs. In the case of the reoviruses, the switch from transcription to replication appears to involve the synthesis of a replicase that converts the (+)mRNAs synthesized early in infection to the double-stranded genome segments.
Viral RNA polymerases, like DNA polymerases, synthesize chains in only one direc-tion; however, in general, RNA polymerases can initiate the synthesis of new chains with-out primers. Thus, there is no obvious end problem in RNA replication. There is one exception to this general rule. The picornaviruses contain a protein that is covalently at-tached to the 5’ end of the genome, called Vpg. This protein is present on the viral RNA because it is involved in the priming of new RNA viral genomes during the infection, similar to the process described earlier for adenoviruses.
Related Topics