Chapter: Biotechnology Applying the Genetic Revolution: Nanobiotechnology
Controlled Change of Protein Shape by DNA
CONTROLLED
CHANGE OF PROTEIN SHAPE BY DNA
Allosteric proteins change
shape in response to the binding of signal molecules (allosteric effectors) at
a specific site. The essence of allosteric control is that the shape change is
transmitted through the protein and affects the conformation of distant regions
of the protein. In allosteric enzymes, binding of an allosteric effector at a
distant site alters the conformation of the active site and may change its
affinity for the substrate. In this way, some enzymes are switched on and off
in response to signal molecules. For example, phosphofructokinase is switched
on by the buildup of AMP, which signals that energy is in short supply. The
response increases flow into the glycolytic pathway. Similarly, many DNA
binding proteins, such as repressors and activators, also change shape on
binding small signal molecules.
It is possible to change the shape of a protein artificially by mechanical force. This has been demonstrated by attaching a single-stranded 60-base segment of DNA between the two poles of a protein. Attaching the DNA requires chemical “handles.” These are engineered into the target protein by replacing amino acids at appropriate positions with cysteine. The reactive SH group is then used to chemically attach the DNA.
Double-helical DNA is much
more rigid than ssDNA. Consequently, the addition of the complementary strand
generates tension as it binds and creates a double helix. This approach has so
far been demonstrated in the laboratory of Giovanni Zocchi at UCLA with maltose
binding protein and an enzyme, guanylate kinase. When maltose binding protein
is stretched, its binding site for maltose opens wider than optimum and the
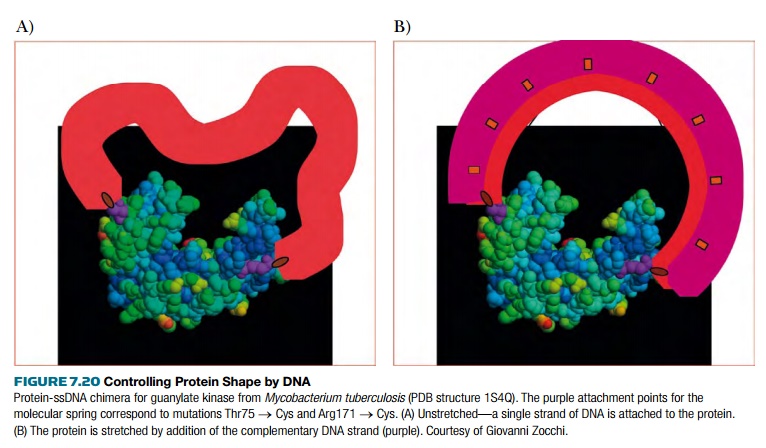
affinity for the sugar decreases. For guanylate kinase (Fig. 7.20), applying
tension decreases enzyme activity by lowering affinity for substrate binding.
In this case releasing the tension by adding DNase to digest the DNA switches the enzyme on
again.
Potential applications are a
long way in the future. However, it is possible to imagine biosensors that
detect DNA sequences based on this mechanism. In addition, it might be possible
to externally control enzymes or other proteins by adding appropriate ssDNA
(or, of course, RNA).
Related Topics