Chapter: Environmental Biotechnology: Genetic Manipulation
Cloning vectors - Basic Principles of Genetic Engineering
Cloning vectors
A cloning vector is frequently a plasmid or a bacteriophage
(bacterial virus) which must be fairly small and fully sequenced, able to
replicate itself when reintroduced into a host cell, thus producing large
amounts of the recombinant DNA for further manipulation. Also it must carry on
it ‘selector marker’ genes. These are different from the reporter genes described
below which are indicators of genomic integrity and activity. A common design
of a cloning vector is one which carries two genes coding for antibiotic
resistance. The ‘foreign’ gene is inserted within one of the genes so that it
is no longer functional therefore it is possible to discriminate by standard
microbiology techniques which bacteria are carrying plasmids containing
recombinant DNA and which are not. Selector genes may operate on at least two
levels, the first at the level of the bacterium, usually Eschericia coli, in which the manipulations are being performed
described aboveand the second being at the level of the final product, for
example, a higher plant. In this case such a selector gene can be resistance to
antibiotics like kanamycin or hygromycin.
Standard cloning vectors
normally carry only selector marker genes required for plasmid construction. To
make the manipulations easier, these genes nor-mally contain a multicloning
site (MCS) which is a cluster of sites for restriction enzymes constructed in
such a way to preserve the function of the gene. Disrup-tion by cloning into
any one of these sites will lose the function of that gene and hence, for
example, if it codes for antibiotic resistance, will no longer protect the
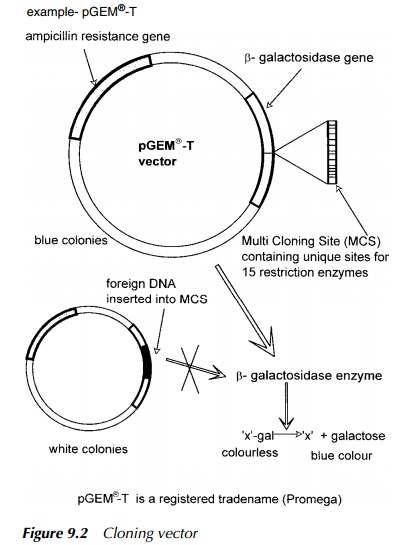
bacterium from that antibiotic. An example is shown in Figure 9.2. This is pGEM
(Promega 1996) which has a MCS in the ß-gal
gene. This codes for ß-galactosidase
from the E. coli lac operon, which
has the capacity to hydrolyse x-gal, a colourless liquid, to produce free
galactose and ‘x’ which results in a blue pigment to the colony. Thus the
screening for successful insertion into the MCS is a simple scoring of blue
(negative) or white (possibly positive) colonies. The success of the experiment
can be determined quickly as this cloning vector also has sequences at either
side of the MCS which allows for rapid DNA sequencing.
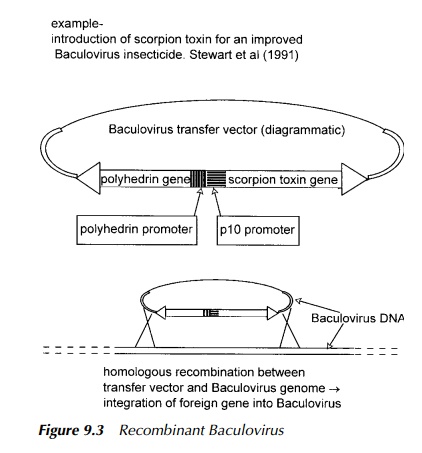
Additionally, some
eukaryotic viruses may be used as vectors but these tend to be so large that
direct cloning into them is difficult. A solution to this is to carry out
manipulations on the desired DNA fragment cloned into a bacterial plasmid and
then transfer the engineered piece into the virus thus making a recombinant
eukaryotic virus. One such virus now used extensively in genetic engineering,
both as a cloning vehicle and as an excellent expression vector, is Baculovirus
illustrated in Figure 9.3 which is also effective as a bioinsecticide.
Related Topics