Chapter: Genetics and Molecular Biology: Nucleic Acid and Chromosome Structure
Topological Considerations in DNA Structure
Topological Considerations in DNA Structure
Topology
introduces a structural feature in addition to the base-paired and helical
aspects of DNA. The origin of this structure can best be understood by
considering a mathematical property of two closed rings. The number of times
that one ring encircles or links the other must be an integral number. It
cannot be changed without physically opening
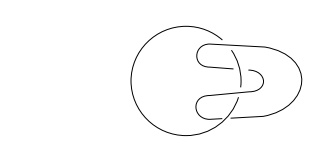
one of
the rings. That is, their linking number is a topological invariant. Many types
of DNA molecules found in cells are covalently closed circles because each
strand is circular. Hence the concept of a linking number applies to DNA
molecules obtained from many sources. The concept also applies to linear DNA if
the ends are prevented from freely rotating, either because of the extreme
length of the DNA or because the DNA is attached to something else.
The
forces tending to hold double-stranded DNA in a right-handed helix with about
10.5 base pairs per turn add a dimension to the analysis of the structures of
covalently closed circles. These forces are suffi-ciently great that the
linking number, Lk, generally
resolves itself into two easily distinguished components: the twist, Tw, which in DNA’s usual right-helical
form has a value of 1 per each 10.5 base pairs, and the writhe, Wr. Twist is the local wrapping of one
of the two strands about the other. If Lk
does not equal Tw, then the
discrepancy must be made up from a global writhing of the molecule as such
global effects can alter the actual number of times one strand encircles the
other. These global effects are called supercoiling or superhelical turns.
Their computation is most difficult because the entire path of the DNA duplex
must be considered. To repeat, for any covalently closed double-stranded DNA
molecule, no matter how it is distorted, unless its phos-phodiester backbone is
broken, Lk = Tw + Wr. This equation
sometimes is written τ = α + β where τ = Lk, α = Tw, and β = Wr. It is curious that the topological invariant Lk equals the sum of two terms, each of
which is not invariant.
It is
convenient to normalize the deviation of the linking number from the normal,
unconstrained value, Lk0.
Since the normal linking is 1 per 10.5 base pairs of DNA, linking values other
than this must change either the twist, the writhe, or both. Rather than just
give the linking number deviation for a DNA molecule, it is more informative to
give the linking number deviation per unit length of the DNA, Lk - Lk0 divided by Lk0 . This value is denoted by σ. Typical values of σ in DNA extracted from bacterial cells are -0.02 to -0.06. Colloquially σ is called supercoiling density, but it must be
remembered that part of the linking number deviation goes into changing the
twist of the DNA molecules and part goes into generating writhe.
Related Topics