Chapter: Medical Microbiology: An Introduction to Infectious Diseases: Bacterial Genetics
Genetic Exchange in Bacteria
GENETIC EXCHANGE
Mutation and selection are important factors in bacterial evolution, but evolution proceeds far faster than it could by these processes alone. For instance, the probability that the process of random mutation alone can produce a cell that, let us say, requires five mutations for optimal growth in a new environment is extremely low. It is in fact the product of the in-dividual mutation frequencies (eg, 10 -6 x10 -6 x 10 -6 x 10 -6 x 10 -6 =10 -30), and that essentially precludes a natural population from ever acquiring the new property in this man-ner. However, such alterations occur because organisms exchange genetic material, thereby permitting combinations of mutations to be collected in individual cells.
Despite the fact that bacteria reproduce exclusively asexually, the sharing of genetic information within and between related species is now recognized to be quite common and to occur in at least three fundamentally different ways. All three processes involve a one-way transfer of DNA from a donor cell to a recipient cell. The molecule of DNA in-troduced into the recipient is called the exogenote to distinguish it from the cell’s own original chromosome, called the endogenote.
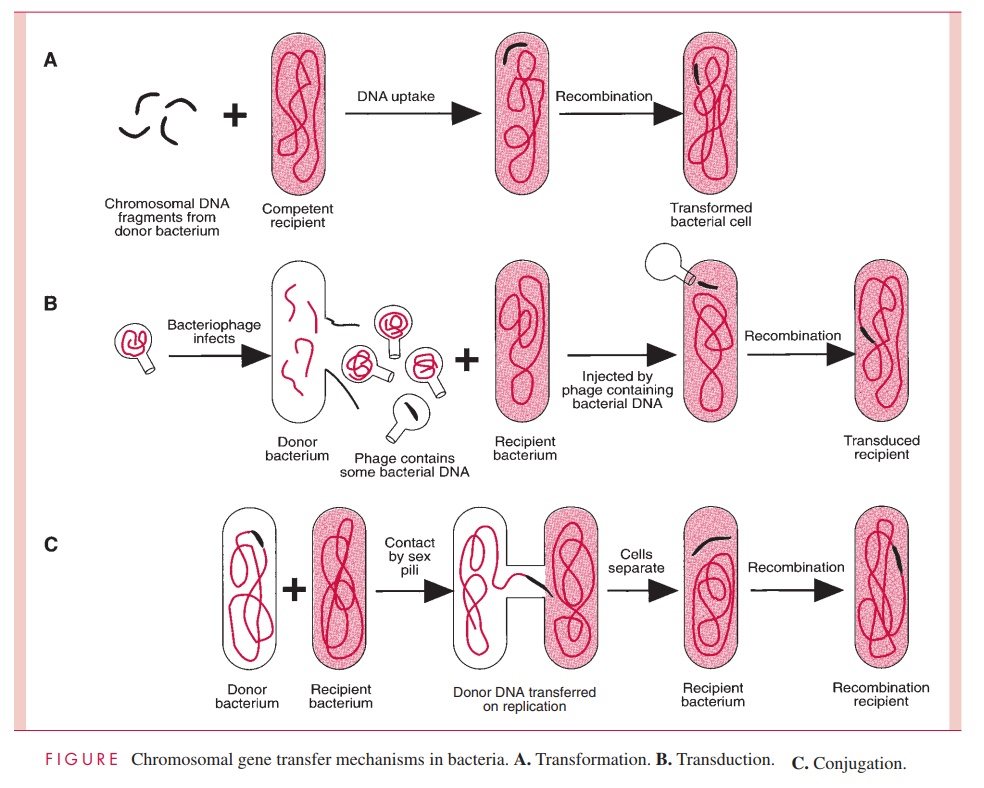
One process of DNA transfer, called transformation, involves the release of DNA into the environment by the lysis of some cells, followed by the direct uptake of that DNA by the recipient cells. By another means of transfer, called transduction, the DNA is introduced into the recipient cell by a nonlethal virus that has grown on the donor cell. The third process, called conjugation, involves actual contact between donor and recipi-ent cell during which DNA is transferred as part of a plasmid (an autonomously replicat-ing, extrachromosomal molecule of circular double-stranded DNA); in conjugation, donor and recipient cells are referred to as male and female, respectively. The three means of gene transfer are summarized in Figure 4 – 2.
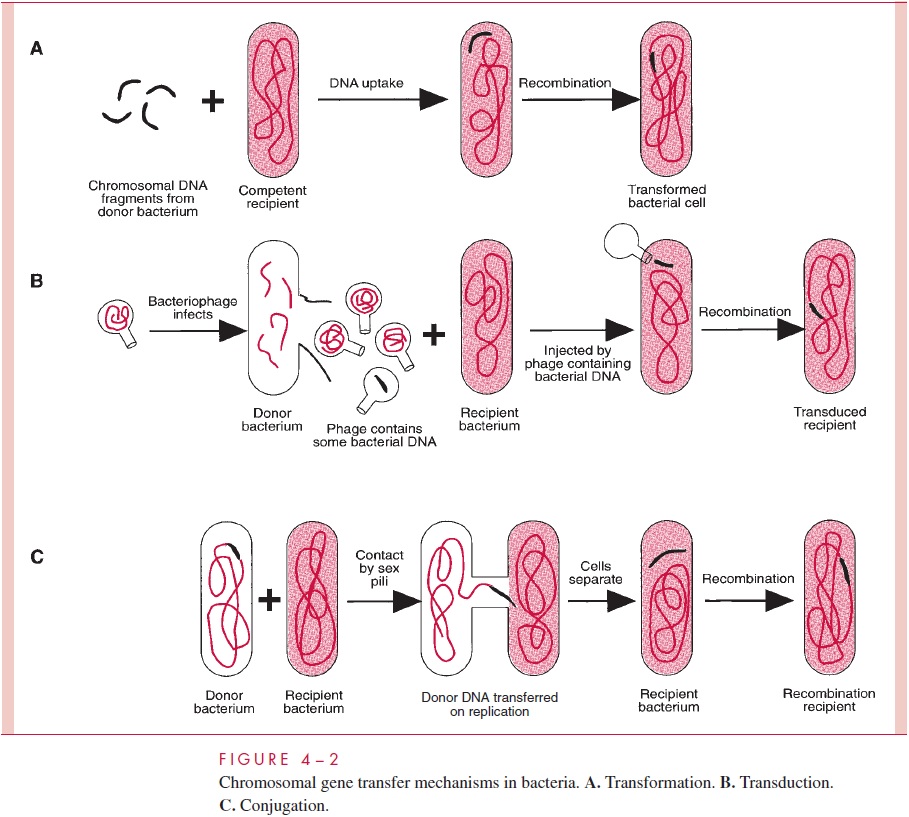
Species of bacteria differ in their ability to transfer DNA, but all three mechanisms are distributed among both Gram-positive and Gram-negative species; however, only transformation is governed by bacterial chromosomal genes. Transduction is totally medi-ated by virus genes, and conjugation, by plasmid genes.
Transformation
Transformation was first demonstrated in 1928 by F. Griffith, a British public health officer, who showed that virulent, encapsulated Streptococcus pneumoniae (pneumococci) that had been killed by heat could confer on living, avirulent, nonencapsulated pneumo-cocci the ability to make the polysaccharide capsule of the killed organisms and thus become virulent for mice. Subsequent work in 1944 by O. T. Avery, C. M. MacLeod, and M. McCarty at the Rockefeller Institute revealed that the “transforming factor” from the dead pneumococci was nothing other than DNA. This discovery had enormous impact on biology, because it was the first rigorous demonstration that DNA is the macromolecule in which genetic information is encoded. It opened the door to modern molecular genetics.
The ability to take up DNA from the environment is called competence, and in many species of bacteria, it is encoded by chromosomal genes that become active under certain environmental conditions. In such species, transformation can occur readily and is said to be natural. Other species cannot enter the competent state but can be made permeable to DNA by treatment with agents that damage the cell envelope making an artificial trans-formation possible.
Natural transformation must be important in nature, judged by the variety of mecha-nisms that different bacteria have evolved to accomplish it. Two of the best-studied systems are those of the Gram-positive pneumococcus and a Gram-negative rod, Haemophilus in-fluenzae. Pneumococcal cells secrete a proteincompetence factorthat induces many of thecells of a culture to synthesize special proteins necessary for transformation, including an autolysin that exposes a cell membrane DNA-binding protein. Any DNA present in the medium is bound indiscriminately; even salmon sperm DNA can be bound and taken up as readily as DNA from another pneumococcal cell.
The surface-bound double-stranded DNA is cleaved into fragments of about 6 to 8 kilobases (kb). One strand is degraded by a nucle-ase, while the complementary strand of each fragment is taken up by a process that seems to be driven by the proton-motive force of the cell membrane . The fate of the internalized DNA fragment then depends on whether it shares homology (the same or simi-lar in base sequence) with a portion of the recipient cell’s DNA. If so, recombination can occur by a process described later, but heterologous DNA (no similarity to the endogenote) is degraded and causes no heritable change in the recipient.
Transformation in H. influenzae is somewhat different. There is no competence factor, and cells become competent merely by growth in an environment rich in nutrients. Only homologous DNA (ie, DNA from the same or a closely related species of Haemophilus) is taken up, and it is taken up in double-stranded form. The selectivity is brought about by the presence of a special membrane protein that binds to an 11-base pair (bp) sequence (5'-AAGTGCGGTCA-3') that occurs frequently in Haemophilus DNA and infrequently in other DNAs. Following binding to molecules of this protein, the homologous DNA is internalized by a mechanism that resembles membrane invagination, resulting in the tem-porary residence of the exogenote in cytosolic membrane vesicles. Although the DNA taken up is double stranded, only one of the two strands participates in the subsequent re-combination with the endogenote.
The common use of E. coli as a host cell in which to clone genes on hybrid plasmids (see Invertible DNA Segments and Recombinational Regulation of Gene Expression) depends on procedures involving treatment with salt and temperature shocks to bring about artificial transformation; this organism has no natural competence mechanism. In contrast, the pathogen, Neisseria gonorrhoeae regularly uses transformation to bring about changes in the antigenic nature of its pili, as described later in the section on recombination.
Transduction
Transduction is virus-mediated transfer of genetic information from donor to recipient cell. To understand transduction and its several mechanisms, it is necessary to preview the nature of bacterial viruses.
Viruses are capable of reproduction only inside living cells. Those that grow in bacte-ria are called bacteriophages, or simply phages. They are minimally composed of pro-tein and nucleic acid, although some may have a very complex structure and composition. The individual virus particle or virion consists of a protein capsid enclosing genomic nu-cleic acid, which is either RNA or DNA, but never both. Virions infect sensitive cells by adsorbing to specific receptors on the cell surface and then, in the case of phages, inject-ing their DNA or RNA. Phages come in two functional varieties according to what hap-pens after injection of the viral nucleic acid. Virulent (lytic) phages cause lysis of the host bacterium as a culmination of the synthesis of many new virions within the infected cell. Temperate phages may initiate a lytic growth process of this sort or can enter a qui-escent form (called a prophage), in which the infected host cell is permitted to proceed about its business of growth and division but passes on to its descendants a prophage genome capable of being induced to produce phage in a process nearly identical to the growth of lytic phages. The bacterial cell that harbors a latent prophage is said to be a lysogen (capable of producing lytic phages), and its condition is referred to as lysogeny.Lysogens are immune to infection by virions of the type they harbor as prophage. Occa-sionally, lysogens are spontaneously induced and lysed by the phage and release mature virions (as many as 75 to 150 or more per cell) into the environment. When triggered by UV irradiation or certain chemicals, an entire population of lysogens are induced simulta-neously to initiate reproduction of their latent virus followed by lysis of the host cells. In-fection of a sensitive cell with the temperate phage can lead to either lysis or lysogeny.
The prophage of different temperate phages exists in one of two different states. In the first, the prophage DNA is physically integrated into a bacterial chromosome; in the sec-ond, it remains separate from the chromosome as an independently replicating, circular-ized, molecule of DNA. Prophages of this sort are in fact plasmids.
For the most part, transduction is mediated by temperate phage, and the two broad types of transduction result from the different physical forms of prophage and the differ-ent means by which the transducing virion is formed. These are termed generalizedtransduction, by which any bacterial gene stands an equal chance of being transduced toa recipient cell, and specialized or restricted transduction, by which only a few genes can be transduced.
Generalized Transduction
Some phages package DNA into their capsids in a nonspecific way, the headful mecha-nism, in which any DNA can be stuffed into the capsid head until it is full. (The head is the principal structure of the virion to which, in some cases, a tail is attached;) An endonuclease then trims off any projecting excess. If fragments of host cell DNA are around during the assembly of mature virions, they can become packaged in place of virus DNA, resulting in pseudovirions. Pseudovirions are the transducing agents. They can ad-sorb to sensitive cells and inject the DNA they contain as though it were viral DNA. The result is the introduction of donor DNA into the recipient cell.
Any given gene has an equal probability of being transduced by this process. With the temperate phage P1 of E. coli, this probability is approximately one transduction event per 105 to 108 virions, because nearly 1 out of every 1000 phage particles made in a P1 lytic infection are pseudovirions, and the bacterial DNA fragments packaged are 1 to 2% of the length of the chromosome. Cotransduction of two bacterial genes by a single pseudovirion occurs only if they are located close together within this small length of the chromosome, and this fact facilitates mapping the position of a newly discovered gene.
Once injected into the host cell, the transduced DNA is lost by degradation unless it can recombine with the chromosome of the recipient cell, usually by homologous recom-bination (see below, Invertible DNA Segments and Recombinational Regulation of Gene Expression) in which both strands of the exogenote cross into and replace the homolo-gous segment of the recipient’s chromosome. However, sometimes the exogenote can per-sist without degradation by assuming a stable circular configuration.
Specialized Transduction
It has been noted that the prophage of some phages is integrated into the lysogen’s chro-mosome. This integration does not occur haphazardly but is restricted to usually one site, called the att (attachment) site. When a lysogen carrying such a prophage is induced to produce virions, excision of the viral genome from the bacterial chromosome occasion-ally (eg, in 1 of 105 to 106 lysogens) occurs imprecisely, resulting in a pickup of genes of the bacterium adjacent to the att site. The resulting virion may be infectious (if no essen-tial phage genes are missing) or defective (if one or more essential genes are missing). In either case, adsorption to a sensitive cell and injection of the DNA can occur, and integra-tion of the aberrant phage genome into the chromosome of the new host cell results in the formation of a lysogen containing a few genes that have been transduced as hitchhikers with the phage genome. Integration of the phage genome automatically accomplishes the recombinational event needed to guarantee reproduction of the transduced genes. Only genes that border the att site stand a chance of being transduced by this process, which is why it is called specialized or restricted transduction.
Because the original pickup event is rare, the first transducing process is termed low-frequency transduction; however, when a lysogenic transductant is, in turn, induced toproduce phage, all of the new virions carry the originally transduced bacterial gene. The resulting mixture of lysed cells and virions now brings about high-frequency transduc-tion of the attached genes.
Bacterial geneticists have learned to move genes of interest near the phage integration site and thereby construct specialized transducing phages containing these genes. Such transducing phages are valuable aids to cloning and sequencing genes and to studying their function and regulation. Obviously a temperate phage that could form a prophage by integrating randomly at any site in the bacterial chromosome would be of special use. The temperate phage Mu of E. coli has this property.
Although both generalized transduction and specialized transduction can be regarded as the result of errors in phage production, transfer of genes between bacterial cells by phage is a reasonably common phenomenon. It occurs at significant frequency in nature; for example, genes conferring antimicrobic resistance in staphylococci are often trans-duced from strain to strain in this way. The toxins responsible for the severe clinical symptoms of diphtheria and of cholera are encoded by genes transduced into Corynebac-terium diphtheriae and Vibrio cholerae, respectively. Transduction is also used exten-sively as a tool in molecular biology research.
Conjugation
Conjugation is the transfer of genetic information from donor to recipient bacterial cell in a process that requires intimate cell contact; it has been likened to mating. By themselves, bacteria cannot conjugate. Only when a bacterial cell contains a self-transmissible plasmid (see below for definition) does DNA transfer occurs. In most cases, conjugationinvolves transfer only of plasmid DNA; transfer of chromosomal DNA is a rarer event, and is mediated by only a few plasmids. Plasmids are of enormous importance to medical microbiology. They are discussed in detail later, but to understand conjuga-tion we should introduce some of their features at this point.
Plasmids are autonomous extrachromosomal elements composed of circular double-stranded DNA; a few rare linear examples have been found. Plasmids are found in most species of Gram-positive and Gram-negative bacteria in most environments. Plasmids gov-ern their own replication by means of special sequences and proteins. They replicate within the host cell (and only within the host cell) and are partitioned between the daughter cells at the time of cell division. In addition, many plasmids are able to bring about their own transfer from one cell to another by the products of a group of genes called tra (for transfer); such plasmids are called conjugative plasmids. Other plasmids, called noncon-jugative, lack this ability. Thetragenes, of which there may be dozens, encode the struc-tures and enzymes that accomplish conjugation. One of these structures is a specialized pilus called the sex pilus, which confers the ability to seize recipient cells on the plasmid-containing donor cells. Retraction of the pilus draws the donor and recipi-ent cell into the intimate contact needed to form a conjugal bridge through which DNA can pass. One strand of the plasmid DNA is then enzymatically cleaved at a site called the ori-gin of transfer (oriT), and the resulting 5’ end of the strand is guided into the recipientcell by the action of various tra-encoded proteins (Fig 4 – 3). Both the introduced strand and the strand remaining behind in the donor cell direct the synthesis of their complemen-tary strand in a process called transfer replication, resulting in complete copies in both donor and recipient cell. Finally, circularization of the double-stranded molecules occurs, the conjugation bridge is broken, and both cells can now function as donor cells.
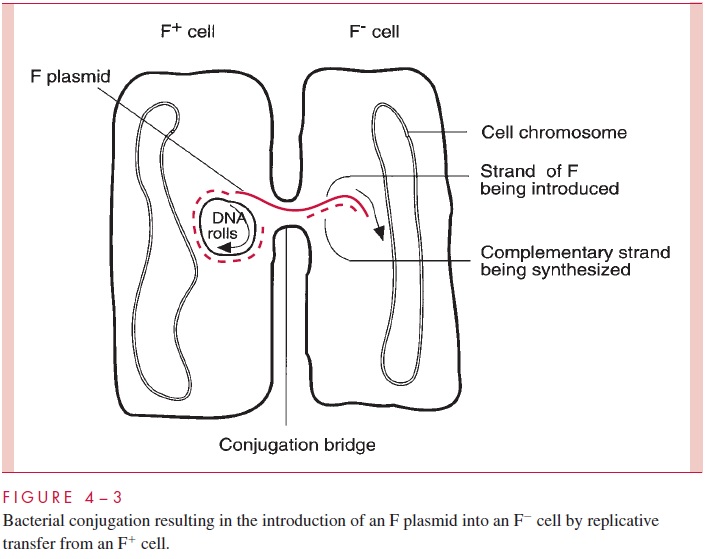
Conjugation is a highly evolved and efficient process. Suitable mixtures of donor and recipient cells can lead to nearly complete conversion of all the recipients into donor, plasmid-containing cells. Furthermore, although some conjugative plasmids can transfer themselves only between cells of the same or closely related species, others are quite promiscuous, promoting conjugation across a wide variety of (usually Gram-negative) species. Conjugation appears to be a carefully regulated process, normally kept in check by the production of a repressor encoded by one of the tra genes. Interestingly, noncon-jugative plasmids that happen to inhabit a cell with a conjugative plasmid can under some circumstances be transferred due to the conjugation apparatus of the latter; this process is called plasmid mobilization. As the later discussion of plasmids shows, their conjugal properties have enormous implications in medicine.
Conjugation in Gram-Negative Species
After many inconclusive attempts by microbiologists to learn whether a sexual process of ge-netic exchange existed among bacteria, J. Lederberg and E. Tatum discovered conjugation in 1946. What they observed was a transfer of chromosomal genes between cells of two differ-ent strains of E. coli. Their discovery stimulated an intensive analysis of the mechanism, leading to the discovery of an agent, the F factor (for fertility factor), that conferred on cells the ability to transfer bacterial chromosome genes to recipient cells. Now it is recognized that the F factor is a conjugative plasmid, although an atypical one in several respects.
The F plasmid is a normal conjugative plasmid in that it possesses many tra genes en-coding a sex pilus (the F-pilus) as well as the ability to form a conjugation bridge, to initi-ate transfer replication, and to perform all the other steps of plasmid transfer. Thus, a cell harboring the F plasmid (an F+cell) can conjugate with a recipient F- cell, and in the process the latter becomes F+ . The process is immediate and efficient because the F factor has lost autoregulation of the conjugation process. However, these properties do not explain how the F plasmid can bring about transfer of chromosomal genes, which is more closely related to another property of F — its ability to integrate at low frequency into the bacterial chromosome at seven or eight chromosomal sites, resulting in linearization of the plasmid DNA as part of the giant circular chromosomal molecule. A cell in which this integration event has occurred is designated a high-frequency recombination (Hfr) cell; it is only this spontaneous mutant in an F- population that transfers donor chromosomal genes. When an Hfr cell encounters an F- cell, conjugation occurs and the usual transfer replication is initi-ated at oriT, within the linear F segment. However, in this circumstance, breaking the integrated plasmid DNA at oriT results in the formation of a linear strand in which the entire bacterial chromosome lies between two portions of the F genome (Fig 4 – 4), and therefore the leading segment of F enters the F cell followed by bacterial genes one after the other. The conjugation bridge usually ruptures long before the entire bacterial chromo-some can be introduced, resulting in the transfer of only one part of the F genome and a variable length of the bacterial chromosome. Thus, conjugation between an Hfr and an F- cell leaves the recipient still F- , but having received bacterial genes; the donor remains Hfr because it retains a copy of the chromosome with its integrated F genome. There are other fertility plasmids, but F remains the best studied.
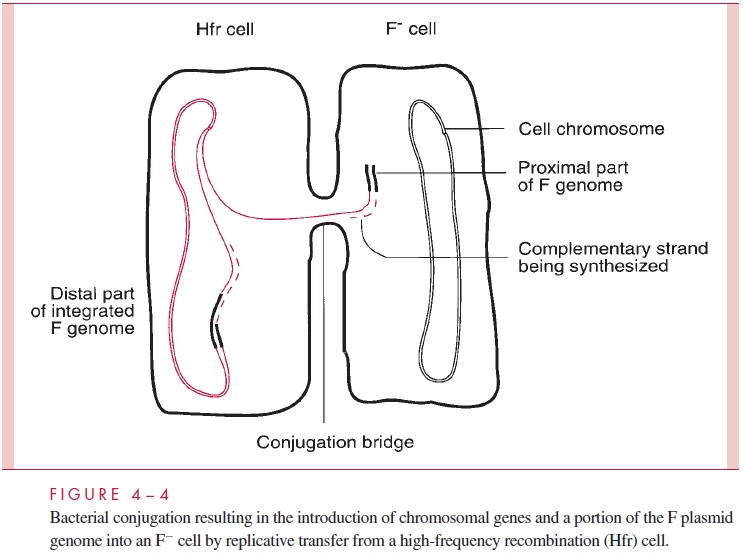
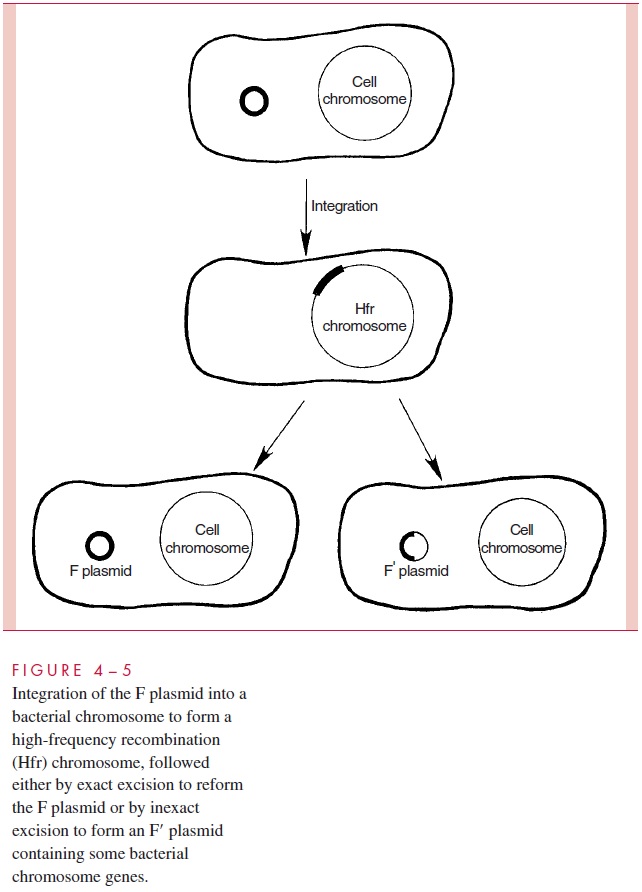
There is an additional wrinkle to the transfer of chromosomal genes by conjugation in E. coli. It is a process termedsexduction,in which an F plasmid transfers from one cellto another a few bacterial chromosomal genes that it happens to contain. This comes about because the F genome in an Hfr cell can, at low frequency, excise itself from the chromosome and circularize into plasmid form. When this excision is imperfect, or in-volves recombinations with other insertion sites, segments of the bacterial chromosome can become included in the plasmid (Fig 4 – 5). When the resulting plasmid, called F to note its content of some bacterial DNA, is transmitted to recipient cells at high frequency by conjugation, the chromosomal genes are transferred as hitchhikers; this is the process of sexduction. By similar processes, segments of bacterial chromosomes can become in-corporated into other plasmids, discussed, that confer resistance to antimicrobics.
Conjugation in Gram-Positive Species
Plasmids carrying genes encoding antimicrobic resistance, common pili and other adhesins, and some exotoxins are readily transferred by conjugation among Gram-positive bacteria in the natural environment as well as in the laboratory. However, conjugation involving chro-mosomal genes may differ between Gram-negative and Gram-positive species, as judged by its characteristics in two well-studied examples, E. coli and Enterococcus faecalis.Conjuga-tion in E. faecalis is mediated by plasmids, but there is also an involvement of chromosomal genes in the process. Donor and recipient cells do not couple by means of a sex pilus but rather by the clumping of cells that contain a plasmid with those that do not. This clumping is the result of interaction between a proteinaceous adhesin on the surface of the donor (plasmid-containing) cell and a receptor on the surface of the recipient (plasmid-lacking) cell. Both types of cells make the receptor (possibly cell wall lipoteichoic acid), but only the plasmid-containing cell can make the adhesin, presumably because it is encoded by a plas-mid gene. Interestingly, donor cells make the adhesin only when in the vicinity of recipient cells because the recipients secrete small peptide pheromones that serve to notify the donor cells of the presence of recipients. Donor cells promptly make adhesin when they sense the pheromone. As a result, clumps are formed, and plasmid DNA is transferred across conju-gation bridges into the recipient cells held in the clumps.
In addition to enterococcal species, species of Bacillus, Staphylococcus, and Clostrid-ium have been found to contain conjugative plasmids. Conjugative transfer of genes hasalso been observed in a number of Gram-positive species in the apparent absence of plasmid DNA. In several instances these transfers involve conjugative transposons (to be discussed later), and it appears that a plasmid intermediate is formed, al-though only transiently.
Before continuing with our discussion of plasmids, we should complete the story of what happens to DNA introduced into recipient cells by any of the three transfer processes, transformation, transduction, and conjugation.
Related Topics