Chapter: Essential Microbiology: Viruses
Replication cycles in animal viruses
Replication cycles in animal viruses
Viruses that infect multicellular organisms such as animals may be specific not only to a particular organism, but also to a particular cell or tissue type. This is known as the tissue tropism of the virus, and is due to the fact that attachment occurs via specificreceptors on the host cell surface.
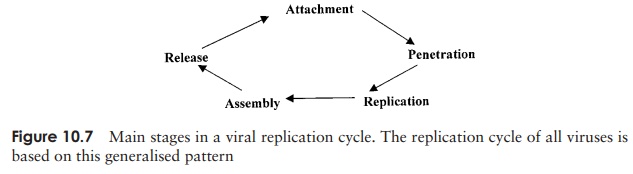
The growth cycles of animal viruses have the same main stages as described for bacteriophages (Figure 10.7), but may differ a good deal in some of the details. Most of these variations are a reflection of differences in structure between bacterial and animal host cells.
Adsorption and penetration
Animal viruses do not have the head and tail structure of phages, so it follows that their method of attachment is different. The specific interaction with a host receptor is made via some component of the capsid, or, in the case of enveloped viruses, by special structures such as spikes (peplomers). Viral attachment sites can frequently be blocked by host antibody molecules; however some viruses (e.g. the rhinoviruses) have overcome this by having their sites situated in deep depressions, inaccessible to the antibodies.
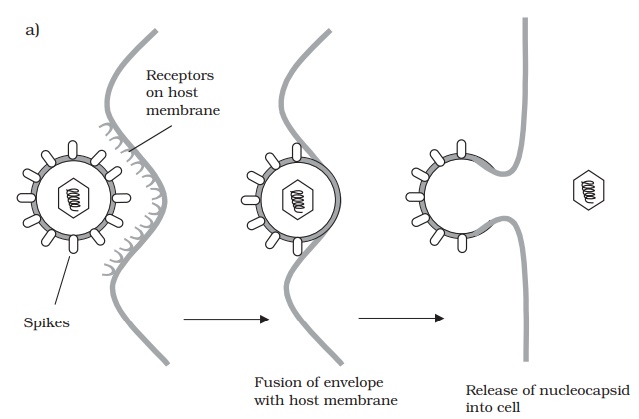
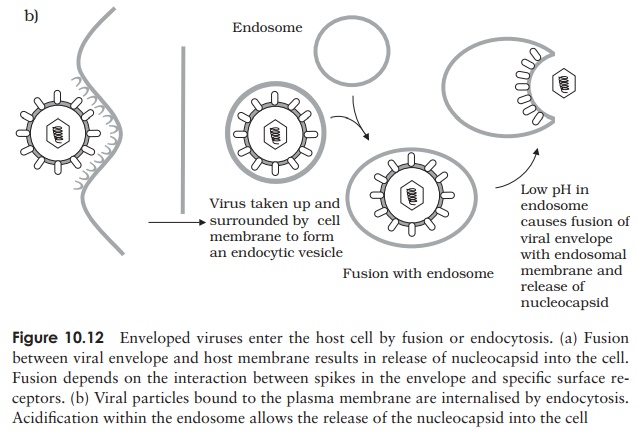
Whereas bacteriophages inject their nucleic acid component from the outside, the process in animal viruses is more complex, a fact reflected in the time taken for com-pletion of the process. Animal viruses do not have to cope with a thick cell wall, and in many such cases the entire virionis internalised. This necessitates the extra step of uncoating, a process carried out by host enzymes. Many animal viruses possess an en-velope; such viruses are taken into the cell either by fusion with the cell membrane, or by endocytosis (Figure 10.12). While some non-enveloped types release only their nu-cleic acid component into the cytoplasm, others require additionally that virus-encoded enzymes be introduced to ensure successful replication.
Replication (DNA viruses)
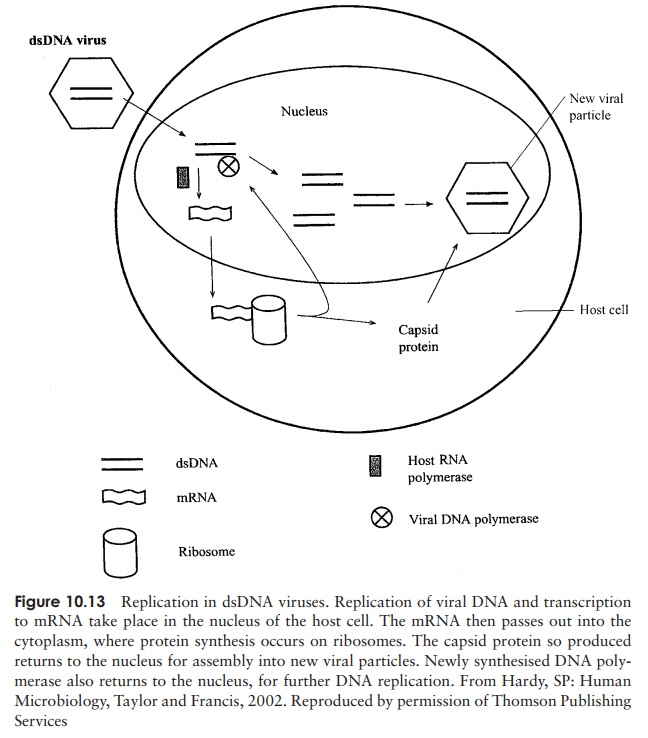
The DNA of animal cells, unlike that of bacteria, is compartmentalised within a nucleus, and it is here that replication and transcription of viral DNA generally occur∗ . Messenger RNA then passes to ribosomes in the cytoplasm for translation (Figure 10.13). In the case of viruses with a ssDNA genome, a double-stranded intermediate is formed, which serves as a template for mRNA synthesis.
Assembly
Translation products are finally returned to the nucleus for assembly into new virus particles.
Release
Naked (non-enveloped) viruses are generally released by lysis of the host cell. In the case of enveloped forms, release is more gradual. The host’s plasma membrane is modified by the insertion of virus-encoded proteins, before engulfing the virus particle and releasing it by a process of budding. This can be seen as essentially the reverse of the process of internalisation by fusion (Figure 10.12a).
Related Topics