Chapter: Basic Concept of Biotechnology : Biotechnology in Forensic Sciences
History of Forensic Genetic Typing
History of Forensic Genetic Typing
More than two decades ago, genetically, polymorphic protein markers were applied to potentially distinguish individuals. But, numerous factors restricted the forensic use of these protein-based genetic systems. The refining power of these markers was low, resultantly, individualization was not difficult. Additionally, the proteins are not available at satisfactory levels for typing in most of the tissues, and they are comparatively more prone to degradation in biological samples in contact with environmental fluctuations.
The typing of genetic polymorphisms at the DNA level assists to overcome these restrictions to a much greater extent. Primarily, there is a remarkable amount of dissimilarities at the DNA level to exploit for individuality testing. Second, any biological substance that comprises nucleated cells potentially can be used for DNA polymorphism typing. Third, DNA is found to be more stable in forensic samples. Consequently, with the current techniques for DNA typing and the set of existing genetic markers, human polymorphism typing at the DNA level is more delicate, more precise, and more instructive than the conventional protein genetic markers. Additionally, DNA technology provides the forensic scientist the greatest possibility to ignore individuals who havebeen falsely connected with a biological sample and to decrease the number of individuals hypothetically incorporated as contributors of the sample to a limited (if not one) individuals.
The early method used regularly for human identity testing (from the mid-1980s) in the US and many other countries was restriction fragment length polymorphism (RFLP) typing of VNTR (variable number of tandem repeat) loci. Typing VNTR loci by RFLP investigation delivered a very high power of judgement, but required samples should hold minimum 10–25 ng of DNA that need to be some what integrated with fragment lengths up to 10,000 bp for any successful typing. Gradually, RFLP analysis was substituted by amplification of DNA (Fig.1) loci by the polymerase chain reaction (PCR) followed by typing of genetic markers. PCR facilitates the scrutinization of samples containing pictogram intensities of DNA, thus enhancing the preciseness of detection one to two folds of magnitude. In fact, small quantities of DNA extracted from different types of samples are typed customarily and effectively using PCR-based assays in forensic laboratories. Sample materials comprise saliva, blood, sweat and semen dropped on different substrates including cigarettes, clothing, postage stamps, drinking straws and containers, envelope flaps, face masks and chewing gum; various tissues from human remains; vaginal swabs from a rape victim; and possible reference samples taken from personal items, such as tooth brushes, hair brushes and razors, which may be advantageous in the identification of anonymous remains.
The initial genetic marker systems examined using PCR-based systems were centred on SNP variation. These SNP-centred systems offered high sensitivity of exposure but did not deliver the power of discernment that VNTR/RFLP typing afforded, and were not very useful for revealing the role of mixed samples because of their limited polymorphism.
To intensify the influence of discrimination of PCR-centred systems, VNTR loci were amplified by polymerase chain reaction, with the allelic forms fragmented by electrophoresis, and successively perceived by silver staining. Short tandem repeat (STR) or microsatellite loci, part of a subclass of highly polymorphic VNTR loci, have substituted the PCR-centred genetic markers. Nowadays, they are used in forensic laboratories world wide sincethey have polymorphic nature, small product size and semi-automated analytical methods. These repeat sequence loci are abundant in the human genome and are highly polymorphic. Autosomal STR loci are comprised of tandemly repeated sequences, each of which is made up of only four to five BP in length. The variable number of alleles of the forensically applicable STR loci ranges usually from 5 to 20 common alleles. The regions covering the repeats are normally small in size and are therefore responsive to amplification by PCR. Because the product length is generally less than 350 bp, much smaller than the length of a DNA fragment prerequisite for VNTR analysis by RFLP, slightly degraded samples are currently typeable. To lessen sample consumption and reduce laboratory manipulations, the STR loci are studied in a multiplex fashion, with approx 15 STR loci amplified and typed at the same time.
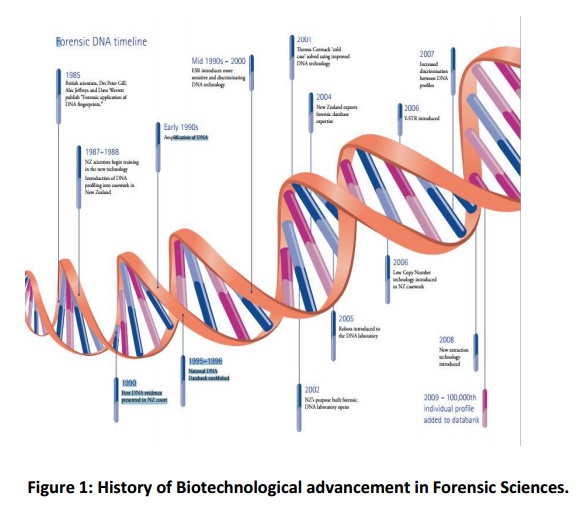
The detailed history of Forensic science is summarised hereunder:
· 1686 –Anatomy Professor Marcello Malpighi mentions in his treaties the ridges, spirals and loops in fingerprints.
· 1839 – H. Baynardprints the first reliable techniques for the detection of sperm, and examines the various microscopic characteristics of various different substrate fabrics.
· 1879 –Rudolph Virchow, a German pathologist, is one of the first to study about hair
· 1892 – Sir Francis Galton in his book “Fingerprints” establishes the individuality of fingerprints and a first classification system for it.
· 1892 – Juan Vucetich develops the fingerprint classification system that was used in Latin America.
· 1901 – Karl Landsteiner notices human blood group.
· 1915 – Prof. Leone Lattes develops the antibody test for ABO blood types.
· 1931 – Franz Josef Holzercomes up with the absorption-inhibition AB typing techniques that were generally used in forensic laboratories.
· 1940 – Rh blood groups were first described by Landsteiner and A. S. Weiner.
· 1953 – James Watson and Francis Crick identify the structure of DNA. 1984 – Sir Alec Jeffreys discovers a method of identifying individuals by DNA –RFLP.
· 1986 – Kerry Mullis discovers PCR method.
· 1986 – DNA is used for the first time to solve a crime by Alec Jeffreys.
· 1987 – DNA profiling is introduced for the first time in a US criminal court.
· 1992 – Thomas Caskey and colleagues suggest the use of short tandem repeats for forensic DNA analysis in their published paper.
· 1996 – Mitochondria DNA evidence is used in US court for the first time.
· 1998 – NIDIS, FBI DNA database, is put into practice.
Related Topics