Chapter: The Diversity of Fishes: Biology, Evolution, and Ecology: Individuals, populations, and assemblages
Genetic structure of populations - Diversity of Fishes
Genetic structure of populations
Determining the physical boundaries of a population can be easy (as in isolated ponds and lakes) or difficult (large rivers and lakes, oceanic regions). Gene flow, the exchange of genes across boundaries and between populations, can blur the distinction between populations. Coregonine whitefishes in many lakes in Canada and sticklebacks in lakes, ponds, and rivers on the Pacific coast of North America are reproductively isolated from each other and consequently form distinct genetic groupings (demes) (Hagen & McPhail 1970; Bell & Richkind 1981; Smith 1981b). In contrast, American Eels, although distributed in ponds, lakes, streams, and rivers from Iceland to Venezuela, all return to a single spawning locale in the Sargasso Sea, remixing their genes at each reproductive episode. Such panmictic spawning means that the entire species consists of only one population (Avise et al. 1986). Similar lack of genetic structure, in terms of genetically discrete populations, has been shown in other anguillids such as European and Japanese Eels (Sang et al. 1994; Lintas et al. 1998),although different authors using different methods have found evidence of genetically unique populations in these species (Wirth & Bernatchez 2001; Tseng et al. 2006). Nearshore marine species generally show levels of gene flow that are strongly related to dispersal capability and distance between populations; good dispersers that are close together are very similar genetically (Waples 1987).
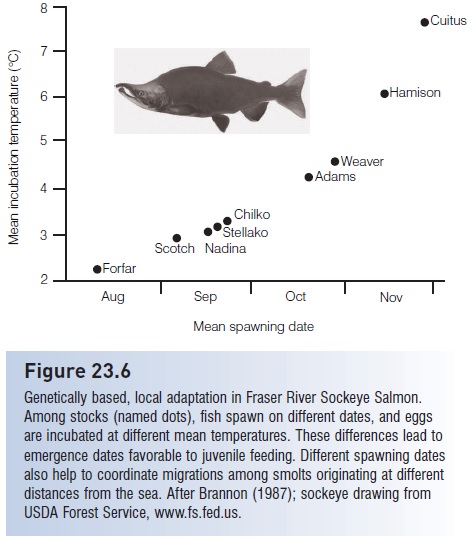
Genetic analysis of high seas species often indicates a lack of differentiation among widely spread populations that were once thought of as several species. Albacore Tuna (Thunnus alalunga) in the North Pacific and South Atlantic and Skipjack Tuna (Katsuwonus pelamis) and Yellowfin Tuna in the Atlantic and Pacific are three examples (Graves & Dizon 1987; Scoles & Graves 1993). Migration over long distances or close proximity in a nearshore marine area do not, however, guarantee gene flow. Salmon of Pacific northwestern rivers (Oncorhynchus spp.) migrate and intermix in the ocean during much of their lives, but genetically discrete stocks separate and return to their natal streams to reproduce, conserving the genetic identity of the more than 200 stocks (Nehlsen et al. 1991; see Fig. 23.6). At the extreme of genetic diversity, rainbow/steelhead trout/salmon, O. mykiss, from the west coast of the Kamchatka Peninsula of Russia differentiate into as many as six distinct life history types (riverine/estuarine, estuarine, anadromous A, anadromous B, anadromous B halfpounders, and resident), with evidence that all six may come from the same spawning redd (Savvaitova et al. 2000; Augerot 2005). Populations of live-bearing Black Surfperch, Embiotoca jacksoni, separated by only 40–80 km in California and Mexico, show genetic differences of a magnitude that is normally found between different species within a genus (Waples 1987; Utter & Ryman 1993).
Even an isolated lake may have more than one genetically distinct population of a species. Arctic Char differentiate into different forms in a number of Scandinavian lakes. In Thingvallavatn, Iceland (the suffi x “vatn” means lake in Icelandic), Arctic Char far outnumber the other two species present, a stickleback and a trout. The Char exist as four distinct forms that occupy different habitats and feed on different food types (see Fig. 24.3). A large and a small morph remain near the bottom and feed on benthic invertebrates in the shallow littoral zone, whereas two other morphs frequent the water column where one feeds on zooplankton in the limnetic (pelagic) region and the other feeds on fishes both inshore and offshore. Morphologically (and appropriately), the benthic-feeding morphs have subterminal mouths and relatively dark coloration, whereas the more pelagic forms have terminal mouths and silvery, countershaded coloration. Spawning times vary among the morphs, and the morphological differences show up shortly after hatching and hence are not ecophenotypic, i.e., they do not result from environmental influences experienced by different individuals.

The morphological and behavioral differences among morphs have a strong genetic basis, as offspring of the different morphs retain their trophic specializations even when raised in a common laboratory environment (Skulason et al. 1993). Genetic differences can also be demonstrated biochemically. Interestingly, the different morphs do not differ genetically, with the exception of the small benthivorous morph, which differs from the other three morphs at one esterase enzyme locus out of 36 loci tested. The morphs may have evolved because of the availability of habitats (adaptive zones or open niches) brought on by the absence of other species in the lake to fill those ecological vacancies (Magnusson & Ferguson 1987; Sandlund et al. 1988). Remarkably, predation by Arctic Char in Thingvallavatn varies suffi ciently across habitats that it has selected for two very different morphs of Threespine Stickleback,Gasterosteus aculeatus (Ólafsdóttir et al. 2007). Sticklebacks are well known for differentiating into distinct benthic and limnetic forms in response to predation pressure and food availability (e.g., McPhail 1994; Schluter 2000). Other fish species demonstrating marked intraspecific diversity, often with a genetic basis, include Crucian Carp, Carassius carassius (Bronmark & Miner 1992);Sockeye Salmon, Oncorhynchus nerka (Hendry & Quinn 1997; Hendry 2001); Pumpkinseed Sunfish, Lepomis gibbosus(Parsons & Robinson 2007); and, of course, Lake Malawi cichlids (Arnegard et al. 1999).
Populations have genetic structure. The distribution of genotypes can be characterized, either by assaying directly for relative frequencies of alleles of genes or by studying the distribution of phenotypes. Genetic analyses of populations can be a powerful technique for solving systematic questions. For example, five species and eight subspecies of kelpfishes (Gibbonsia, Clinidae) were recognized from western North America, based on traditional meristic and morphometric analyses. Reanalysis using allozyme data obtained by electrophoresis of enzymes extracted from various tissues indicated very little genetic differentiation among the subspecies and even between some species, suggesting a high degree of gene flow among populations (Stepien & Rosenblatt 1991). Three nominal species, the Scarlet Kelpfish (G. erythra), the Crevice Kelpfish (G. montereyensis), and an offshore Mexican endemic (G. norae), showed no significant differences inthe frequencies of different alleles at 40 gene loci (the term nominal refers to a population that has been described as a separate species).
Reanalysis of morphometric data showed that many anatomical differences were instead sexual dimorphisms or anatomical trends that changed continually as a function of water temperature or with depth (the latter are ecophenotypic differences because they represent phenotypes that differ consistently and result from depth of occurrence). All scarlet kelpfish, distinguished by more caudal peduncle scales and a higher dorsal spine, were in fact males. All nominally Crevice Kelpfish were females. G. norae differed only in lower counts of scale rows and fin rays, but it is generally observed that fish that develop in warmer water lay down fewer meristic elements (Barlow 1961; Meristic variation). Hence G. erythra, G. norae, and G. montereyensis represent different populations of a single species, G. montereyensis. Subspecies of kelpfishes also showed little genetic differentiation. Anatomical differences occurred in populations that occupied different
depth zones, but characters such as dorsal spine height generally increased with depth of occurrence and were probably ecophenotypic, not genetic, traits. The combined analysis – genetic, morphological, distributional, and ecological – indicated that three rather than five species and no subspecies of kelpfish existed. Hence, genetic analysis showed the true taxonomic relationships among the different species.
Understanding the genetic make-up of a population has become increasingly important as environmental degradation and overexploitation place many populations and species at risk. A key character is the degree of genetic variation in a population. Genetic variation results from selection, mutation, dispersal (emigration out of and gene flow into populations), non-random mating, and genetic drift (random changes in gene frequencies, particularly in small, isolated populations). Genetic variation is a chief driving force of evolution; natural selection acts on such variation, favoring genotypes that are adapted to current conditions. Measures of genotypic variation and frequencies can tell us whether a population has become dangerously inbred and lacks the genetic diversity necessary to allow for adaptation to changing environmental conditions, whether gene flow is occurring between populations, and whether hybridization with introduced species is occurring (and hence if the genetic identity of a species is threatened) (Schonewald-Cox et al. 1983; Meffe 1986).
Recognizing the importance of genetic structure among populations has led to the concept of metapopulations, which describes populations of a species linked by gene flow via migration, recruitment, or colonization. Metapopulations are thought to exist in a variety of species, including widely distributed, commercially exploited marine fishes (e.g., Atlantic Cod, Atlantic Herring; McQuinn 1997; Wright et al. 2006), among fishes in a stream network (Cutthroat Trout, Bayou Darters; Slack et al. 2004; Neville et al. 2006), among anadromous and estuarine species (Pacific Salmon, Tidewater Goby; Policansky & Magnuson 1998; Lafferty et al. 1999), and among populations of reef fishes (surgeonfish, parrotfish; Planes et al. 1996; Geertjeset al. 2004). Such a focus has important implications for the management of fishery species as well as for the design of protected species and areas because it points to linkages among distant locales (e.g., Fausch et al. 2002). For example, protecting crucial habitat and populations in a few specific locales along a 1000 km long region of the Gulf of California (Sea of Cortez) is likely to assure larval sources linked by ocean currents to many other Gulf areas of biological and socioeconomic importance (e.g., Sala et al. 2002).
Related Topics