Chapter: Essential Microbiology: Biochemical Principles
Biomacromolecules
Biomacromolecules
Many of the most important molecules in biological systems are polymers, that is, large molecules made up of smaller subunits joined together by covalent bonds, and in some cases in a specific order.
Carbohydrates
Carbohydrates are made up of just three different ele-ments, carbon, hydrogen and oxygen. The simplest carbohydrates are monosaccharides, or simple sugars; these
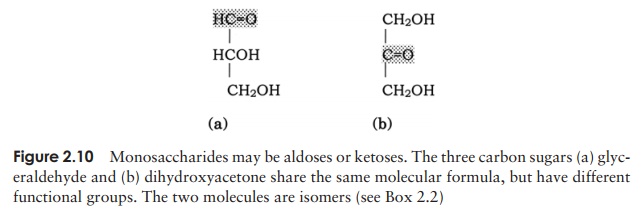
have the general formula (CH2O)n. They are classed as either aldoses or ketoses, ac-cording to whether they contain an aldehyde group or a ketone group (Figure 2.10). Monosaccharides can further be classified on the basis of the number of carbon atoms they contain. The simplest are trioses (three carbons) and the most important biologi-cally are hexoses (six carbons).
Monosaccharides are generally crystalline solids which are soluble in water and have a sweet taste. They all reducing sugars, so called because they are able to reduce alkaline solutions of cupric ions (Cu2+ ) to cuprous ions (Cu+ ).
A disaccharide is formed when two monosaccharides (which may be of the same type or different), join together with a concomitant loss of a water molecule (Fig-ure 2.11). Further monosaccharides can be added, giving chains of three, four, five
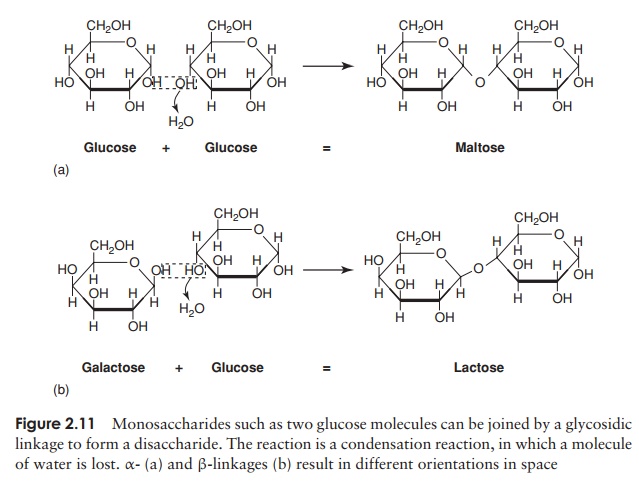
or more units. These are termed oligosaccharides (oligo, a few), and chains with many units are polysaccharides. The chemical bond joining the monosaccharide units to-gether is called a glycosidic linkage. The bond between the two glucose molecules that make up maltose is called an α-glycosidic linkage; in lactose, formed from one glu-cose and one galactose, we have a β-glycosidic linkage. The two bonds are formed in the same way, with the elimination of water, but they have a different orienta-tion in space. Thus disaccharides bound together by α- and β-glycosidic linkages have a different overall shape and as a result the molecules behave differently in cellular metabolism.
Biologically important molecules such as starch, cellulose and glycogen are all polysaccharides. Another is dextran, a sticky substance produced by some bacteria to aid their adhesion. They differ from monosaccharides in being generally insoluble in water, not tasting sweet and not being able to reduce cupric ions. Most polysaccharides
Proteins
Of the macromolecules commonly found in living systems, proteins are the most ver-satile, having a wide range of biological functions and this fact is reflected in their structural diversity.
The five elements found in most naturally occurring proteins are carbon, hydrogen, oxygen, nitrogen and sulphur. In addition, other elements may be essential components of certain specialised proteins such as haemoglobin (iron) and casein (phosphorus).
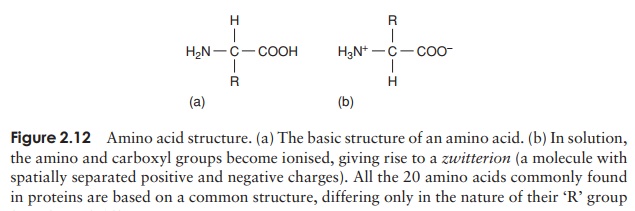
Proteins can be very large molecules, with molecular weights of tens or hundreds of thousands. Whatever their size, and in spite of the diversity referred to above, all proteins are made up of a collection of ‘building bricks’ called amino acids joined together. Amino acids are thought to have been among the first organic molecules formed in the early history of the Earth, and many different types exist in nature. All these, including the 20 commonly found occurring in proteins, are based on a common structure, shown in Figure 2.12. It comprises a central carbon atom (known as the α-carbon) covalently bonded to an amino (NH2) group, a carboxyl (COOH) group and a hydrogen atom. It is the group attached to the final valency bond of the α-carbon which varies from one amino acid to another; this is known as the ‘R’-group.
The 20 amino acids found in proteins can be conveniently divided into five groups, on the basis of the chemical nature of their ‘R’-group. These range from a single hydrogen atom to a variety of quite complex side chains (Figure 2.13). It is unlikely nowadays that you would need to memorise the precise structure of all 20, as the author was asked to do in days gone by, but it would be advisable to familiarise yourself with the groupings and examples from each of them. The groups differentiate on the basis of a polar/non-polar nature and on the presence or absence of an ionisable ‘R’-group.
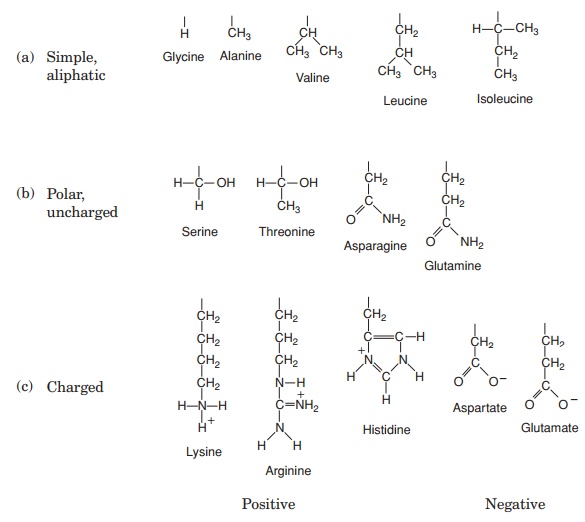
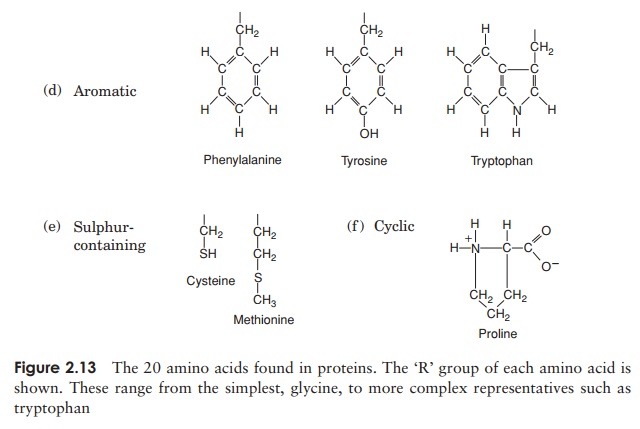
Note that one amino acid, proline, falls outside the main groups. This differs from the others in that it has one of its N—H linkages replaced by an N—C, which forms part of a cyclic structure (Figure 2.13). This puts certain conformational constraints upon proteins containing proline residues.
As can be seen from Figure 2.13, the simplest amino acid is glycine, whose R-group is simply a hydrogen atom. This means that the glycine molecule is symmetrical, with a hydrogen atom on opposite valency bonds. All the other amino acids however, are asymmetrical. The α-carbon acts as what is known as a chiral centre, giving the molecule right or left ‘handedness’. Thus two stereoisomers known as the D- and L-forms are possible for each of the amino acids except glycine. All the amino acids found in naturally occurring proteins have the L-form; the D-form also occurs in nature but only in certain specific, non-protein contexts.
Proteins, as we’ve seen, are polymers of amino acids. Amino acids are joined together by means of a peptide bond. This involves the -NH2 group of one amino acid and the -COOH group of another. The formation of a peptide bind is a form of condensation reaction in which water is lost (Figure 2.14).
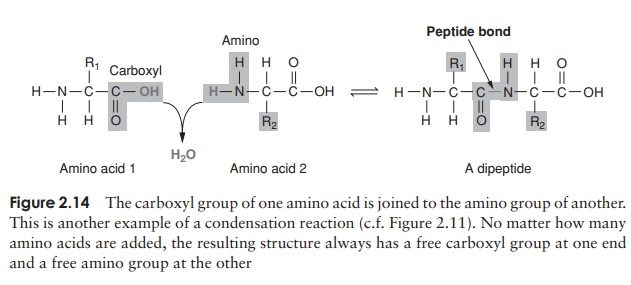
The resulting structure of two linked amino acids is called a dipeptide; note that this structure still retains an -NH2 at one end and a -COOH at the other. If we were to add on another amino acid to form a tripeptide, this would still be so, and if we kept on adding them until we had a polypeptide, we would still have the same two groupings at the extremities of the molecule. These are referred to as the N-terminus and the C-terminus of the polypeptide. Since a water molecule has been removed at the formation of each peptide bond, we refer to the chain so formed as being composed of amino acid residues, rather than amino acids. The actual distinction between a protein and a polypeptide based on the number of amino acid residues is not clear-cut; generally, with over 100, we refer to proteins, but some naturally occurring proteins are a lot smaller than this.
So far, we can think of proteins as long chains of many amino acid residues, rather like a string of beads. This is called the primary structure of the protein; it is deter-mined by the relative proportions of each of the 20 amino acids, and the order in which they are joined together. It is the basis of all the remaining levels of structural complexity, and it ultimately determines the properties of a particular protein. It is also what makes one pro-tein different from another. Since the 20 types of amino acid can be linked together in any order, the number of possible sequences is astronomical, and it is this great variety of structural possibilities that gives proteins such diverse structures and functions.
Some parts of the primary sequence are more important than others. If we took a protein of, say, 200 amino acid residues in length, took it apart and reassembled the amino acids in a different order, we would almost certainly alter (and probably lose completely) the properties of that protein. If we look at the primary sequence of a protein molecule which serves essentially the same function in several species, we find that nature has allowed slight alterations to occur during evolution, but these are often conservative substitutions, where an amino acid has been replaced by a similar one (one from the same group in Figure 2.13), and thus have little effect on the pro-tein’s properties. In certain parts of the primary sequence, such substitutions are less well tolerated, for example the few residues that make up the active site of an enzyme. In cases such as the one above, alterations have not been allowed at these points in the primary sequence, and the sequence is the same, or almost so, in all species possessing that protein. The sequence in question is said to have been conserved.
Higher levels of protein structure
The structure of proteins is a good deal more complicated than a just a linear chain of amino acids. A long thin chain is unlikely to be very stable; proteins therefore undergo a process of folding which makes the molecule more stable and compact. The results of this folding are the secondary and tertiary structures of a protein
The secondary structure is due to hydrogen bonding between a carbonyl (-CO) group and an amido (-NH) group of amino acid residues on the peptide backbone (Figure 2.15). The ‘R’ group plays no part in secondary protein structure. Two regular patterns of folding result from this; the α-helix and the β-pleated sheet.
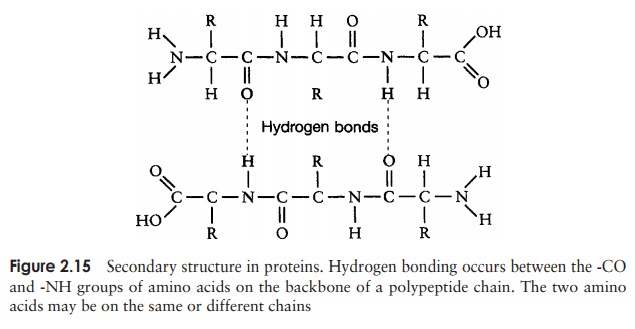
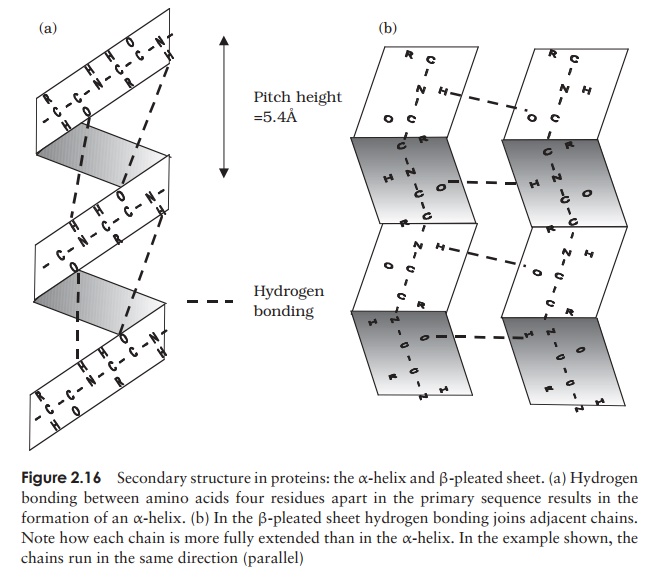
The α-helix occurs when hydrogen bonding takes place between amino acids close together in the primary structure. A stable helix is formed by the -NH group of an amino acid bonding to the -CO group of the amino acid four residues further along the chain (Figure 2.16a). This causes the chain to twist into the characteristic he-lical shape. One turn of the helix occurs every 3.6 amino acid residues, and results in a rise of 5.4 A (0.54 nm); this is called the pitch height of the helix. The ability to form a helix like this is dependent on the component amino acids; if there are too many with large R-groups, or R-groups carrying the same charge, a stable helix will not be formed. Because of its rigid structure, proline (Figure 2.13) cannot be accommodated in anα-helix. Naturally occurring α-helices are always right-handed, that is, the chain of amino acids coils round the central axis in a clockwise direction. This is a much more stable configuration than a left-handed helix, due to the fact that there is less steric hindrance (overlapping of electron clouds) between the R-groups and the C==O group on the peptide backbone. Note that if proteins were made up of the D-form of amino acids, we would have the reverse situation, with a left-handed form favoured. In the β-pleated sheet, the hydrogen bonding occurs between amino acids either on separate polypeptide chains or on residues far apart in the primary structure (Figure 2.16b). The chains in a β-pleated sheet are fully extended, with 3.5 A (0.35 nm) between adjacen amino acid residues (c.f. α-helix, 1.5 A). When two or more of these chains lie next to each other, extensive hydrogen bonding occurs between the chains. Adjacent strands in a β-pleated sheet can either run in the same direction (e.g. N→C), giving rise to a par-allel β-pleated sheet, or in opposite directions (antiparallel β-pleated sheet, as shown in Figure 2.16b).
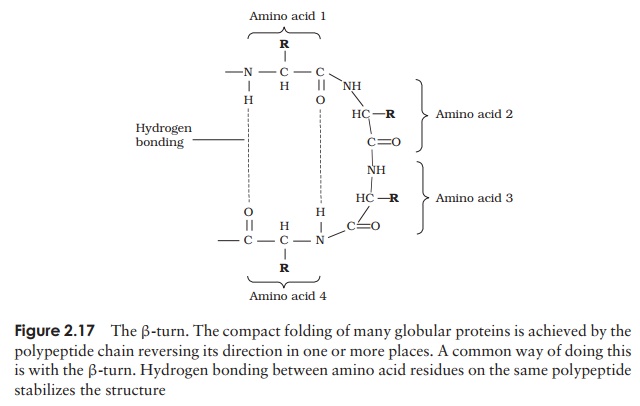
A common structural element in the secondary structure of proteins is the β-turn. This occurs when a chain doubles back on itself, such as in an antiparallel β-pleated sheet. The -CO group of one amino acid is hydrogen bonded to the -NH group of theresidue three further along the chain. Frequently, it is called a hairpin turn, for obvious reasons (Figure 2.17). Numerous changes in direction of the polypeptide chains result in a compact, globular shape to the molecule.
Typically about 50 per cent of a protein’s secondary structure will have an irregular form. Although this is often referred to as random coiling, it is only random in the sense that there is no regular pattern; it still contributes towards the stability of the molecule. The proportions and combinations in which α-helix, β-pleated sheet and random coiling occur varies from one protein to another. Keratin, a structural protein found in skin, horn and feathers, is an example of a protein entirely made up of α-helix, whilst the lectin (sugar-binding protein) concanavalin A is mostly made up of β-pleated sheets.
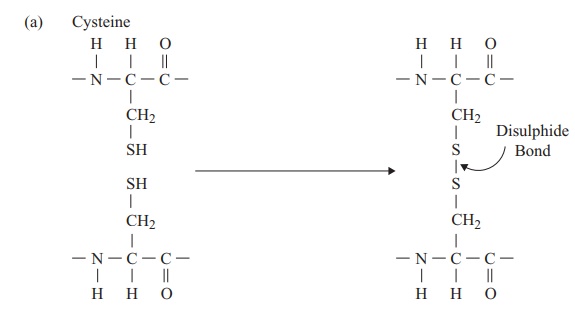
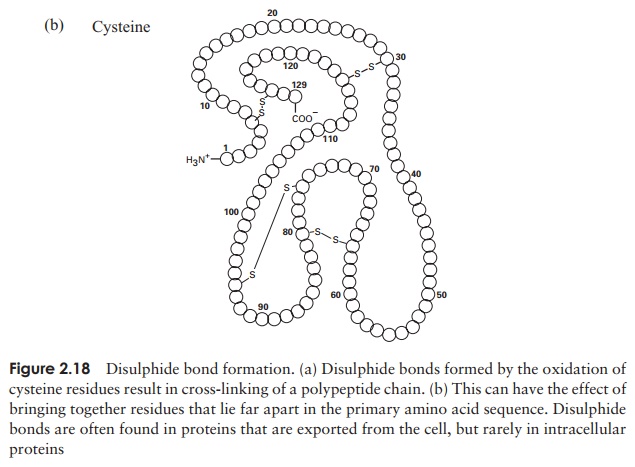
The tertiary structure of a protein is due to interactions between side chains, that is, R-groups of amino acid residues, resulting in the folding of the molecule to produce a thermodynamically more favourable structure. The structure is formed by a variety of weak, non-covalent forces; these include hydrogen bonding, ionic bonds, hydrophobic interactions, and Van der Waals forces. The strength of these forces diminishes with distance, therefore the formation of a compact structure is encouraged. In addition, the -SH groups on separate cysteine residues can form a covalent -S—S- linkage. This is known as a disulphide bridge and may have the effect of bringing together two cysteine residues that were far apart in the primary sequence (Figure 2.18).
In globular proteins, the R-groups are distributed according to their polarities; non-polar residues such as valine and leucine nearly always occur on the inside, away from the aqueous phase, while charged, polar residues including glutamic acid and histidinegenerally occur at the surface, in contact with the water.
The protein can be denatured by heating or treatment with certain chemicals; this causes the tertiary structure to break down and the molecule to unfold, resulting in a loss of the protein’s biological properties. Cooling, or removal of the chemical agents, will lead to a restora-tion of both the tertiary structure and biological activity, showing that both are entirely dependent on the primary sequence of amino acids.
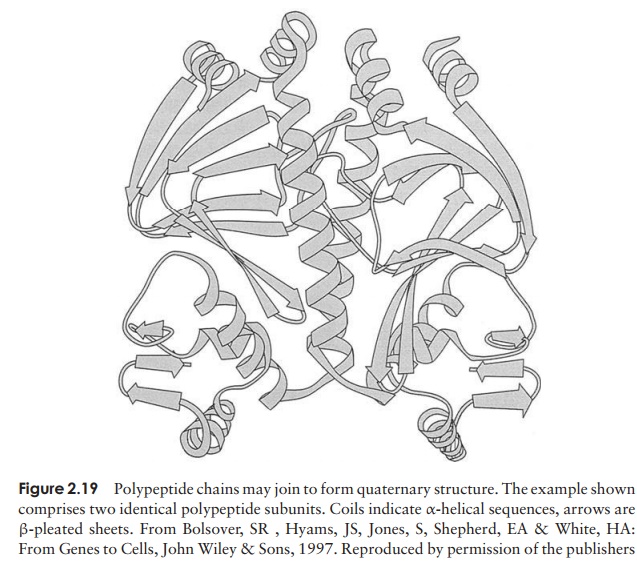
Even the tertiary structure is not always the last levelof organisation of a protein, because some are made up of two or more polypeptide chains, each with its own secondary and tertiary structure, combined together to give the quaternary structure (Figure 2.19). These chains may be identical or different, dependingon the protein. Like the tertiary structure, non-covalent forces between R-groups are responsible, the difference being that this time they link amino acid residues on separate chains rather than on the same one.
Such proteins lose their functional properties if dissociated into their constituent units; the quaternary joining is essential for their activity. Phosphorylase A, an enzyme involved in carbohydrate metabolism, is an example of a protein with a quaternary structure. It has four subunits, which have no catalytic activity unless joined together as a tetramer.
Although all proteins are polymers of amino acids existing in various levels of structural complexity as we have seen above, some have additional, non-amino acid components. They may be organic, such as sugars (gly-coproteins) or lipids (lipoproteins) or inorganic, includ-ing metals (metalloproteins) or phosphate groups (phos-phoproteins). These components, which form an integral part of the protein’s structure, are called prosthetic groups.
Nucleic acids
The third class of polymeric macromolecules are the nucleic acids. These are deoxyri-bonucleic acid (DNA) and ribonucleic acid (RNA), and both are polymers of smaller molecules called nucleotides. As we shall see, there are important differences both in the overall structures of RNA and DNA and in the nucleotides they contain, so we shall consider each of them in turn.
The structure of DNA
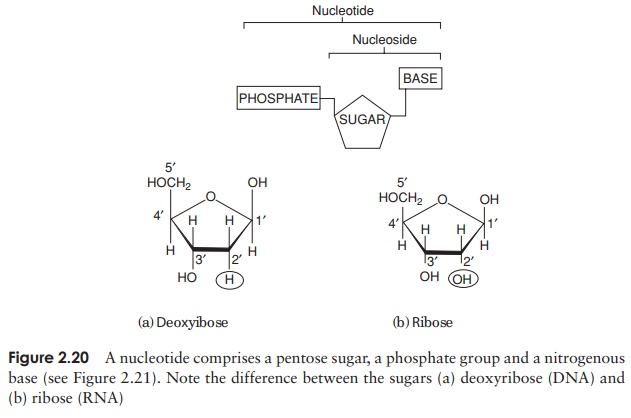
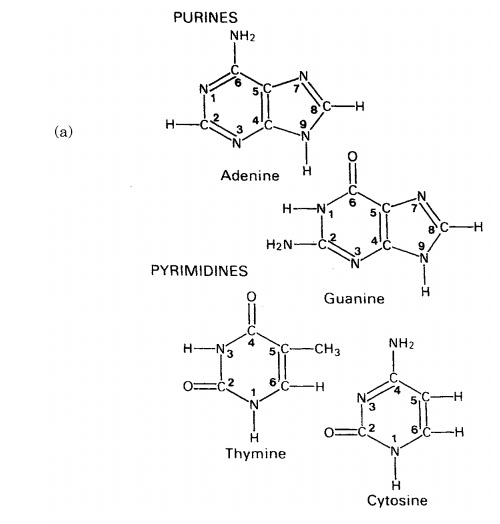
The composition of a DNA nucleotide is shown in Figure 2.20(a). It has three parts, a five-carbon sugar called deoxyribose, a phosphate group and a base. This base can be any one of four molecules; as can be seen in Figure 2.21, these are all based on a cyclic structure containing nitrogen. Two of the bases, cytosine and thymine, have a single ring and are called pyrimidines. The other two, guanine and adenine, have a double ring structure; these are the purines. The four bases are often referred to by their initial letter only, thus we have A, C, G and T.
One nucleotide differs from another by the identity of the base it contains; the rest of the molecule (sugar and phosphate) is identical. You will recall from the previous section that the properties of a protein depend on the order in which its constituent amino acids are linked together; we have exactly the same situation with nucleic acids, except that instead of an ‘alphabet’ of 20 ‘letters’, here we have one of only four. Nevertheless, because nucleic acid molecules are extremely long, and the bases can occur in almost any order, an astronomically large number of different sequences is possible.
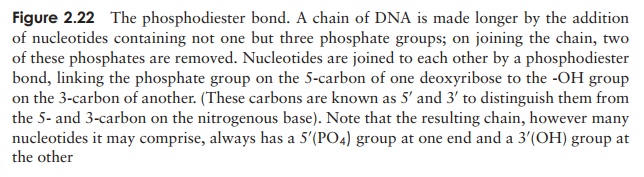
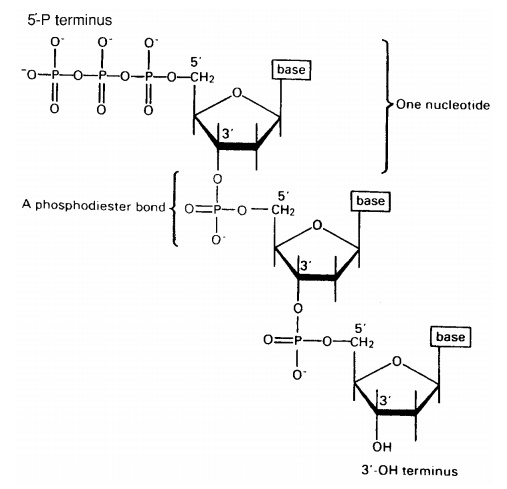
The nucleotides join together by means of a phosphodiester bond. This links the phosphate group of one base to an -OH group on the 3-carbon of the deoxyribose sugar of another (Figure 2.22). The chain of nucleotides therefore has a free -OH group attached to the 3-carbon (the 3 end) and a free phosphate group attached to the 5-carbon (the 5 end). This remains the case however long the chain becomes.

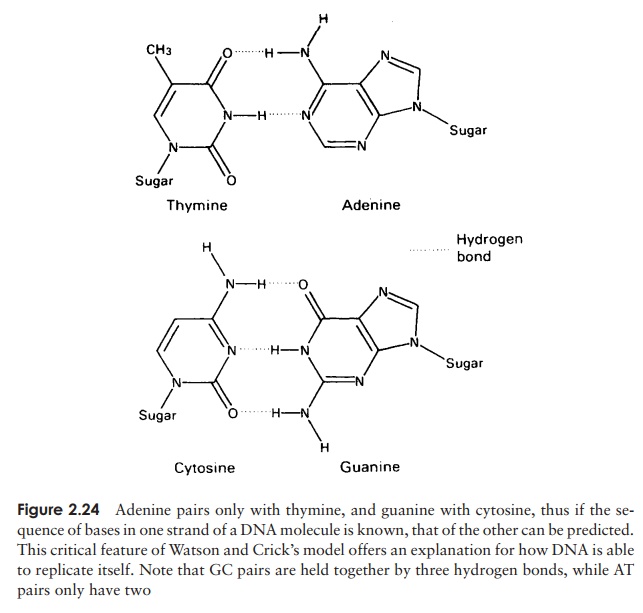
The structure of DNA however is not just a sin-gle chain of linked nucleotides, but two chains wound around each other to give the double helix form made famous by the model of James Watson and Francis Crick in 1953 (Figure 2.23). If we com-pare this to an open spiral staircase, alternate sugar and phosphate groups make up the ‘skeleton’ of the stair-case, while the inward-facing bases pair up by hydro-gen bonding to form the steps. Notice that each nu-cleotide pair always comprises three rings, resulting from a combination of one purine and one pyrimidine base. This means that the two strands of the helix are always evenly spaced. The way in which the bases pair is further governed by the phenomenon of complementary basepairing. A nucleotide containing thymine will only pair with one containing adenine,and likewise guanine always pairs with cytosine (Figure 2.24). Thus, the sequence of
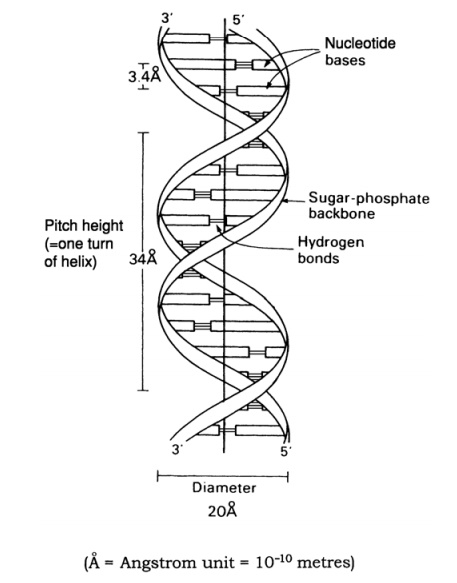

nucleotides on one strand of the double helix determines that of the other, as it has a complementary structure. Figure 2.23 shows how the two strands of the double helix are antiparallel, that is they run in opposite directions, one 5 → 3 and the other 3 →5 . We shall look at how this structure was used to propose a mechanism for the way in which DNA replicates and genetic material is copied.
The structure of RNA
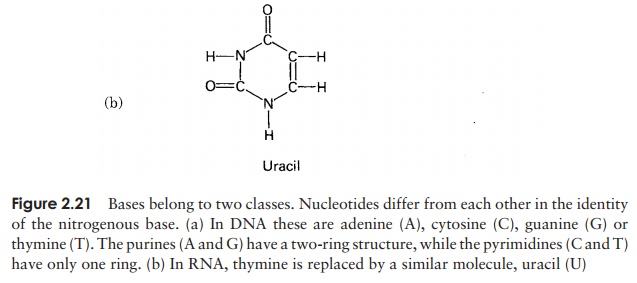
In view of the similarities in the structure of DNA and RNA, we shall confine ourselves here to a consideration of the major differences. There are two important differences in the composition of nucleotides of RNA and DNA. The central sugar molecule is not deoxyribose, but ribose; as shown in Figure 2.20, these differ only in the possession of an -H atom or an -OH group attached to carbon-2. Second, although RNA shares three of DNA’s nitrogenous bases (A, C and G), instead of thymine it has uracil. Like thymine, this pairs specifically with adenine.
The final main difference between RNA and DNA is the fact that RNA generally comprises only a single polynucleotide chain, although this may be subject to secondary and tertiary folding as a result of complementary base pairing within the same strand. The roles of the three different forms of RNA will be discussed.
Lipids
Although lipids can be large molecules, they are not regarded as macromolecules because unlike proteins, polysaccharides and nucleic acids, they are not polymers of a basic subunit. Moreover, lipids do not share any single structural characteristic; they are a diverse group structurally, but have in common the fact that they are insoluble inwater, but soluble in a range of organic solvents. This non-polar nature is due to thepredominance of covalent bonding, mainly between atoms of carbon and hydrogen.
Fats are simple lipids, whose structure is based on fatty acids. Fatty acids are long hydrocarbon chains ending in a carboxyl (-COOH) group. They have the
CH3—(CH2)n—COOH
wheren is usually an even number. They combine with glycerol according to the basic reaction:
Alcohol + Acid → Ester
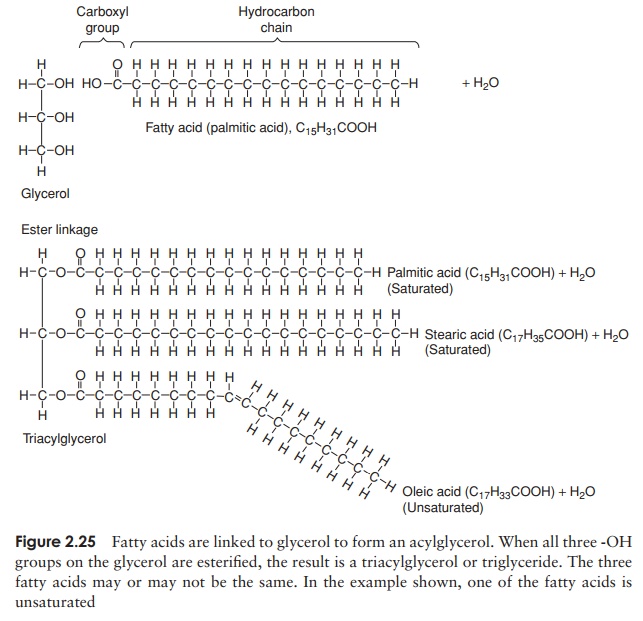
The bond so formed is called an ester linkage, and the result is an acylglycerol (Figure 2.25). One, two or all three of the -OH groups may be esterified with a fatty acid, to give respectively mono-, di- and triacylglycerols. Natural fats generally contain a mixture of two or three different fatty acids substituted at the three positions; consequently, a considerable diversity is possible among fats. Fats serve as energy stores; a higher proportion of C—C and C—H bonds in comparison with proteins or carbohydrates results in a higher energy-storing capacity.
The second main group of lipids to be found in living cells arephospholipids. These have a similar structure to triacylglycerols, except that instead of a third fatty acid chain, they have a phosphate group joined to the glycerol (Figure 2.26), introducing a hydrophilic element to an otherwise hydrophobic molecule. Thus, phospholipids are an example of an amphipathic molecule, with a polar region at one end of the
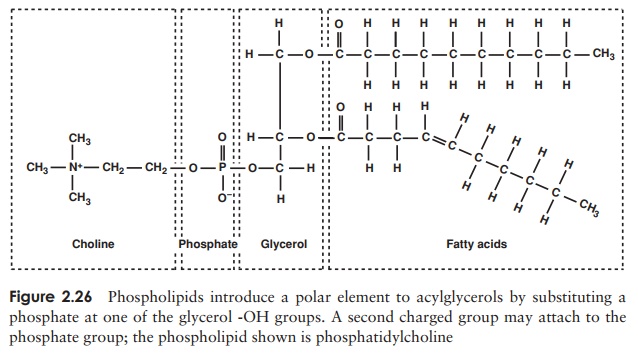

molecule and a non-polar region at the other. This fact is essential for the forma-tion of a bilayer when the phospholipid is introduced into an aqueous environment; the hydrophilic phosphate groups point outwards towards the water, while the hy-drophobic hydrocarbon chains ‘hide’ inside (Figure 2.27, and c.f. Figure 2.8, micelle formation).
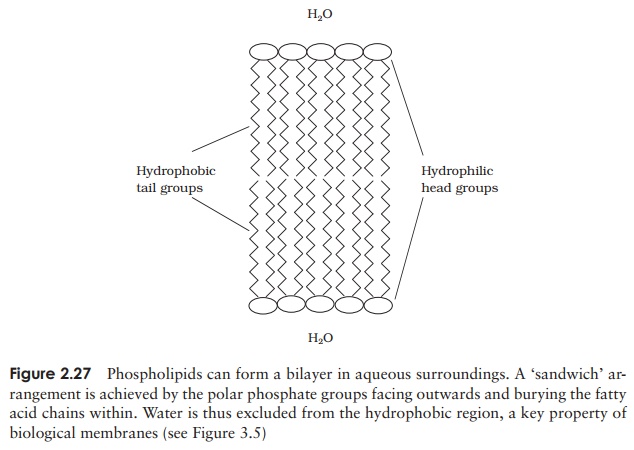
This bilayer structure forms the basis of all biological membranes, forming a barrier around cells and certain organelles. Phospholipids generally have an-other polar group attached to the phosphate; Figure 2.25 shows the effect of substituting serine.
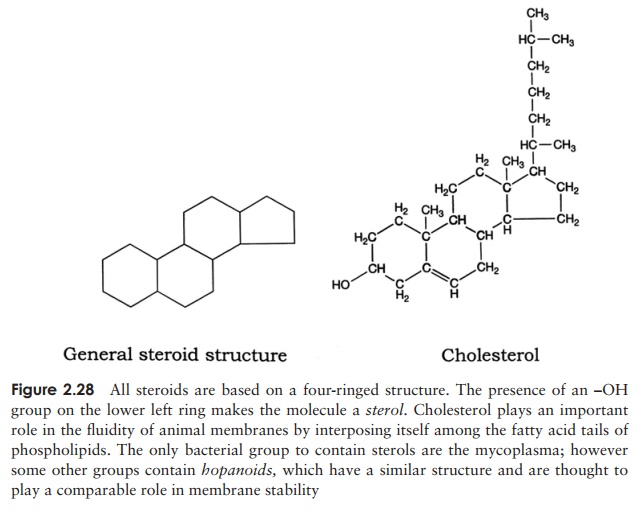
The structural diversity of lipids can be illustrated by comparing fats and phospho-lipids with the final group of lipids we need to consider, the steroids. As can be seen from Figure 2.28, these have a completely different form, but still share in common the property of hydrophobicity. The four ring planar structure is common to all steroids, with the substitution of different side groups producing great differences in function. Cholesterol is an important component of many membranes.
It would be wrong to gain the impression that living cells contain only molecules of the four groups outlined above. Smaller organic molecules play important roles as precursors or intermediates in metabolic pathways, and several inorganic ions such as potassium, sodium and chloride play essential roles in maintaining the living cell. Finally, some macromolecules comprise elements of more than one group, for example, lipopolysaccharides (carbohydrate and lipid) and glycoproteins (protein and carbohydrate).
Related Topics