Chapter: Biochemistry: Transcription of the Genetic Code: The Biosynthesis of RNA
Transcription in Prokaryotes
Transcription in Prokaryotes
RNA Polymerase in Escherichia coli
The
enzyme that synthesizes RNA is called RNA
polymerase, and the most extensively studied one was isolated from E. coli. The molecular weight of this
enzyme is about 470,000 Da, and it has a multisubunit structure. Five different
types of subunits, designated α, ω, β, β', and σ, have been identified. The actual composition
of the enzyme is α2ωββ'σ . The σ-subunit is rather loosely bound to the rest of
the enzyme (the α2ωββ' portion), which is called
the core enzyme. The holoenzyme consists of all the
subunits, including the σ-subunit.
What do the subunits of RNA polymerase do?
The σ-subunit is involved in the
recognition of specific promoters, whereas the β-, β'-, α-, and ω-subunits combine to make the active site for
polymerization.The mechanism of the polymerization reaction is an active area
of research.
Which of the DNA strands is used in transcription?
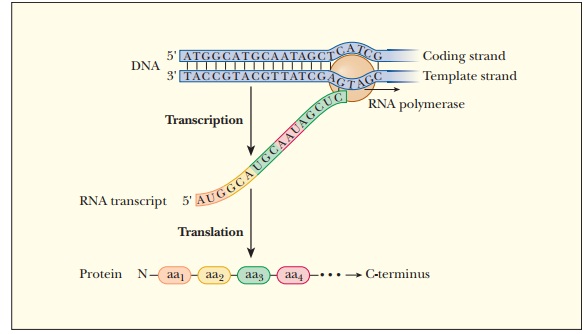
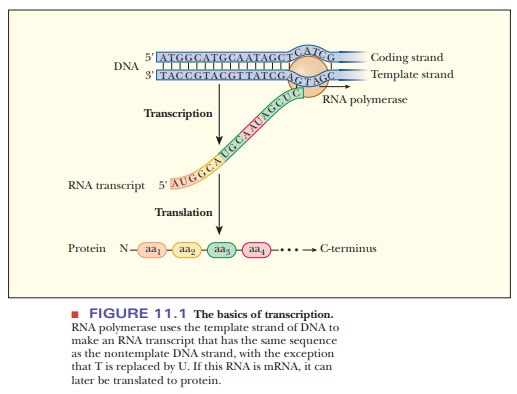
Figure
11.1 shows the basics of information transfer from DNA to protein. Of the two
strands of DNA, one of them is the template for RNA synthesis of a particular
RNA product. RNA polymerase reads it from 3' to 5'. This strand has several
names. The most common is the template
strand, because it is the strand that directs the synthesis of the RNA. It
is also called the antisense strand,
because its code is the complement of the RNA that is produced. It is sometimes
called the (–) strand by convention.
The other strand is called the coding
strand because its sequence of DNA will be the same as the RNA sequence that
is produced (with the exception of U replacing T). It is also called the sensestrand, because the RNA sequence
is the sequence that we use to determinewhat amino acids are produced in the
case of mRNA. It is also called the (+)strand
by convention, or even the
nontemplate strand. For our purposes, wewill use the terms template strand and coding strand throughout. Because the DNA in the coding strand has
the same sequence as the RNA that is produced, it is used when discussing the
sequence of genes for proteins or for promoters and controlling elements on the
DNA.
The core enzyme of RNA polymerase is catalytically active but lacks specific-ity. The core enzyme alone would transcribe both strands of DNA when only one strand contains the information in the gene. The holoenzyme of RNA polymerase binds to specific DNA sequences and transcribes only the correct strand. The essential role of the σ-subunit is recognition of the promoter locus (a DNA sequence that signals the start of RNA transcription;). The loosely bound σ-subunit is released after transcription begins and about 10 nucleotides have been added to the RNA chain. Prokaryotes can have more than one type of σ-subunit.
The nature of the σ-subunit
can direct RNA poly-merases to different promoters and cause the transcription
of various genes to reflect different metabolic conditions.
Promoter Structure
Even the
simplest organisms contain a great deal of DNA that is not transcribed. RNA
polymerase must have a way of knowing which of the two strands is the template
strand, which part of the strand is to be transcribed, and where the first
nucleotide of the gene to be transcribed is located.
How does RNA polymerase know where to begin transcription?
Promoters
are DNA sequences that provide direction for RNA polymerase. The promoter
region to which RNA polymerase binds is closer to the 3' end of the template
strand than is the actual gene for the RNA to be synthesized. The RNA is formed
from the 5' end to the 3' end, so the polymerase moves along the
template strand from the 3' end to the 5' end. However, by convention,
all control sequences are given for the coding strand, which is 5' to 3'. The
binding site for the polymerase is said to lie upstream of the start of transcription, which is farther to the 5' side of
the coding strand. This is often a source of confusion to students, and it is
important to remember the correct orientation. The promoter sequence will be
given based on the coding strand, even though the RNA polymerase is actually
binding to the template strand. Promoters are upstream, which means to the 5' side of
the coding strand and to the 3' side of the template strand.
Most
bacterial promoters have at least three components. Figure 11.2 shows some
typical promoter sequences for E. coli
genes. The component closest to the first nucleotide to be incorporated is
about 10 bases upstream. Also by con-vention, the first base to be incorporated
into the RNA chain is said to be at position +1 and is called the transcription start site (TSS). All the
nucleotides upstream from this start site are given negative numbers.
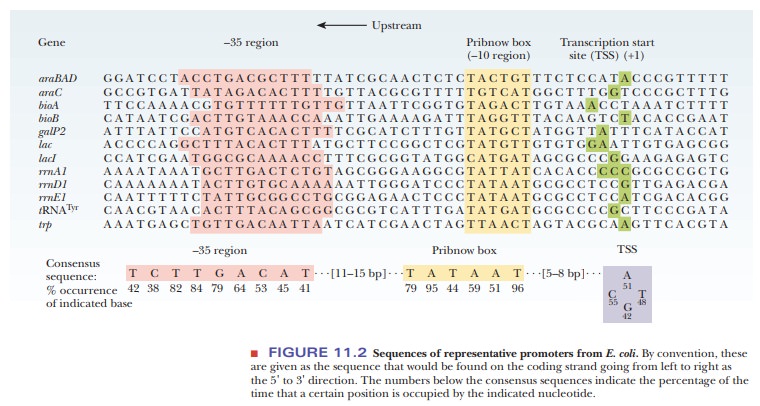
Because the first promoter element is about 10 bases upstream, it is called the –10 region, but is also called the Pribnow box after its discoverer. After the Pribnow box, there are 16 to 18 bases that are completely variable. The next promoter element is about 35 bases upstream of the TSS and is simply called the –35 region or –35 element.
An element is a general
term for a DNA sequence that is somehow important in controlling transcription.
The area from the –35 element to the TSS is called the core promoter. Upstream of the core promoter can be an UP element, which enhances the binding
of RNA polymerase. UP elements usually extend from –40 to –60. The region from
the end of the UP element to the transcription start site is known as the extendedpromoter.
The base
sequence of promoter regions has been determined for a number of prokaryotic genes,
and a striking feature is that they contain many bases in common. These are
called consensus sequences. Promoter
regions are A–T rich, with two hydrogen bonds per base pair; consequently, they
are more easily unwound than G–C-rich regions, which have three hydrogen bonds
per base pair. Figure 11.2 shows the consensus sequences for the –10 and –35
regions.
Even
though the –10 and –35 regions of many genes are similar, there are also some
significant variations that are important to the metabolism of the organism.
Besides directing the RNA polymerase to the correct gene, the pro-moter base
sequence controls the frequency with which the gene is transcribed. Some
promoters are strong, and others are weak. A strong promoter binds RNA
polymerase tightly, and the gene therefore is transcribed more often. In
general, as a promoter sequence varies from the consensus sequence, the
bind-ing of RNA polymerase becomes weaker.
Chain Initiation
The process of transcription (and translation as well, as we will see is usually broken down into phases for easier studying. The first phase of transcription is called chain initiation, and it is the part of transcription that has been studied the most. It is also the part that is the most controlled.
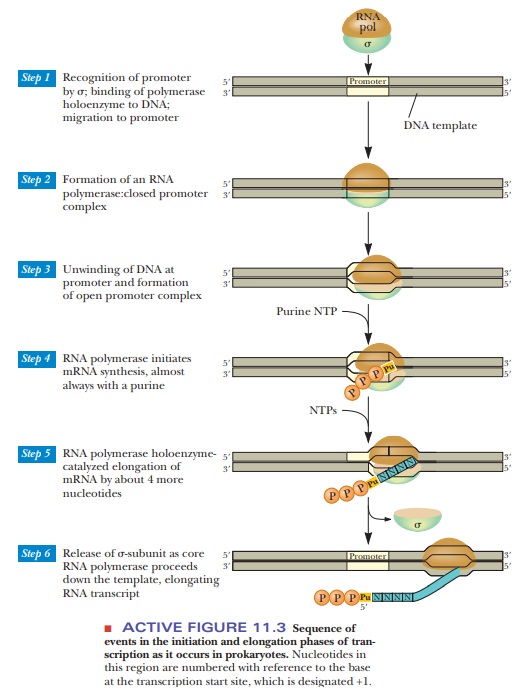
Chain initiation begins when RNA polymerase (RNA pol) binds to the promoter and forms what is called the closed complex (Figure 11.3). The σ-subunit directs the polymerase to the promoter. It bridges the –10 and –35 regions of the promoter to the RNA polymerase core via a flexible “flap” in the σ-subunit.
Core enzymes
lacking the σ-subunit
bind to areas of DNA that lackpromoters. The holoenzyme may bind to
“promoterless” DNA, but it dissoci-ates without transcribing.
Chain
initiation requires formation of the open
complex. Recent studies show that a portion of the β' and the σ-subunits initiate strand separation, melting
about 14 base pairs surrounding the transcription start site. A purine
ribonucleoside triphosphate is the first base in RNA, and it binds to its
comple-mentary DNA base at position +1. Of the purines, A tends to occur more
often than G. This first residue retains its 5'-triphosphate group (indicated
by ppp in Figure 11.3).
Chain Elongation
After the strands have separated, a transcription bubble of about 17 base pairs moves down the DNA sequence to be transcribed (Figure 11.3), and RNA polymerase catalyzes the formation of the phosphodiester bonds between the incorporated ribonucleotides. When about 10 nucleotides have been incorporated, the σ-subunit dissociates and is later recycled to bind to another RNA polymerase core enzyme.
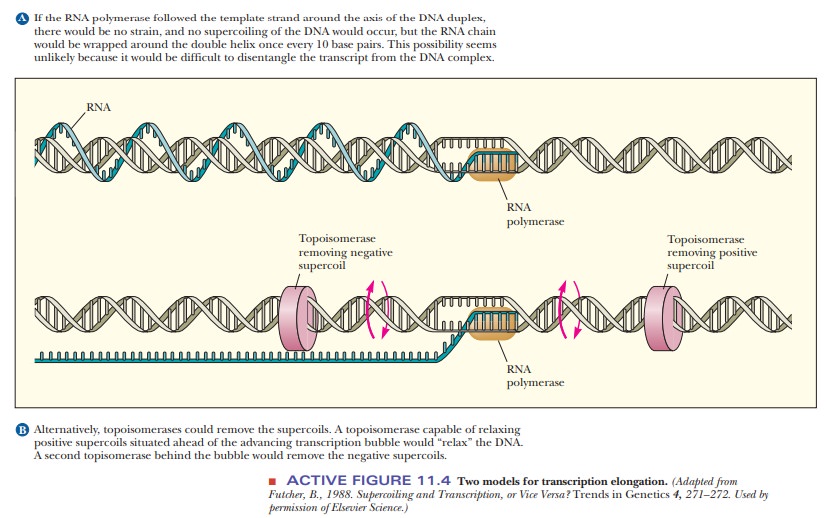
The
transcription process supercoils DNA, with negative supercoiling upstream of
the transcription bubble and positive supercoiling downstream, as shown in
Figure 11.4. Topoisomerases relax the supercoils in front of and behind the
advancing transcription bubble. The rate of chain elongation is not constant.
The RNA polymerase moves quickly through some DNA regions and slowly through
others. It may pause for as long as one minute before continuing.
Instead of finishing every RNA chain, RNA polymerase actually releases most chains near the beginning of the process, after about 5 to 10 nucleotides have been assembled, in a process called abortive transcription. The cause of abortive transcription is the failure of RNA polymerase to break its own bonds to the promoter via the σ-subunit. Studying this process has led to the current model of the mechanism of transcription. In order for chain elongation to occur, the RNA polymerase must be able to launch itself off the promoter. Given the tight binding between the σ-subunit and the promoter, this requires substantial energy. Figure 11.5 shows the current model of the open complex that allows progression into chain elongation. The RNA polymerase is bound tightly to the DNA promoter. It “scrunches” the DNA into itself, causing torsional strain of the separated DNA strands. Like a bow loading up with potential energy as the bowstring is pulled, this provides the energy to allow the polymerase to break free.
Chain Termination
Termination
of RNA transcription also involves specific sequences downstream of the actual gene for the RNA to be transcribed. There
are two types of termination mechanisms. The first is called intrinsic termination, and it is
controlled by specific sequences called termination
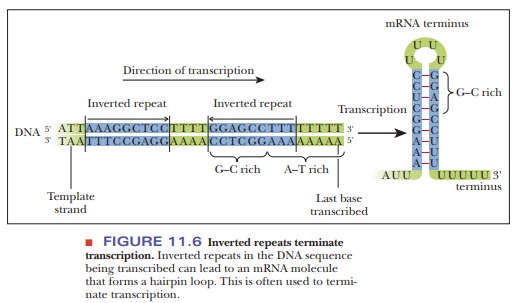
sites. The termination sites are characterized by two inverted repeats
spaced by a few other bases (Figure 11.6). Inverted repeats are sequences of
bases that are complementary, such that they can loop back on themselves. The
DNA then encodes a series of uracils. When the RNA is created, the inverted
repeats form a hairpin loop. This tends to stall the advancement of RNA
polymerase. At the same time, the presence of the uracils causes a series of
A–U base pairs between the template strand and the RNA. A–U pairs are weakly
hydrogen-bonded compared with G–C pairs, and the RNA dissociates from the
transcription bubble, ending transcription.
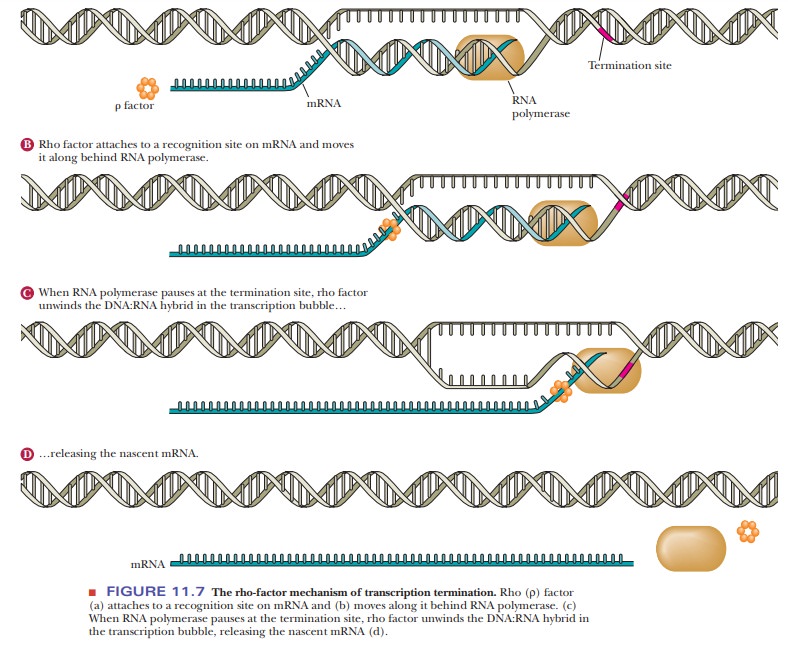
The other type of termination involves a special protein called rho (ρ). Rho-dependent termination sequences also cause a hairpin loop to form. In this case, the ρ protein binds to the RNA and chases the polymerase, as shown in Figure 11.7. When the polymerase transcribes the RNA that forms a hairpin loop (not shown in the figure), it stalls, giving the ρ protein a chance to catch up. When the ρ protein reaches the termination site, it facilitates the dissociation of the transcription machinery. The movement of the ρ protein and the dissociation require ATP.
Summary
Prokaryotic transcription is catalyzed by RNA
polymerase, which is a 470,000-Da enzyme with five types of subunits: α, ω, β, β', and σ.
RNA polymerase moves along the template strand
of DNA and produces a complementary RNA sequence that matches the coding strand
of DNA.
RNA polymerase recognizes
specific DNA sequences called promoters and binds to the DNA. The promoters
tell the polymerase which DNA should be transcribed.
The σ-subunit is loosely bound to RNA polymerase and
is involved in promoter recognition.
Transcription
can be divided into three parts-initiation, elongation, and termination.
Related Topics