Chapter: Genetics and Molecular Biology: Chemotaxis
Methylation and Adaptation
Methylation and Adaptation
The biochemical components in the cell’s rapid
response, adaptation response, and signaling pathways have been found. They are
phospho-rylation and methylation of several of the proteins necessary for
chemo-taxis. First we will discuss methylation. Adler had observed that
methionine is necessary for chemotaxis. That is, a methionine auxot-roph grown
in the presence of methionine would be chemotactic. Upon removal of methionine,
however, the cells remained motile, but could no longer swim up gradients of
attractant. Auxotrophs for other amino acids remained chemotactic upon amino
acid starvation.
The results discussed above indicate that
methionine plays some special role in chemotaxis. Later, when the mechanism of
chemotaxis was understood to be modulation in the frequency of runs and
tumbles, it was sensible to ask what methionine did. Experiments showed that
methionine-starved cells are unable to tumble. The next question was whether
methionine starvation destroyed the physical ability to tumble or whether it
eliminated the signal to tumble. At first this question seems unapproachable.
Genetics, however, came to the rescue. Among the many types of nonchemotactic
mutants known are some that inces-santly tumble. The question then was whether
these tumbling mutants continued to tumble during methionine starvation. All that
was neces-sary to answer this question was to make these tumbling mutants
methionine auxotrophs and to starve them of methionine. After this treatment
they continued to tumble. This shows methionine is not necessary for tumbling,
and therefore that the amino acid must be part of a pathway that signals
tumbling to occur.
How might methionine be required? Methionine via
S-adenosyl methionine is a common source of methyl groups in metabolism. Most
likely, then, the methionine requirement hints that methylation is involved in
the tumbling signal. Consistent with this notion is the fact discussed above
that the addition of arsenate to cells, which blocks ATP formation and hence
S-adenosyl methionine synthesis, also blocks chemotaxis but not motility.
With such clues for the involvement of methylation
in chemotaxis, it is natural to look for methylated proteins. The level of
methylation of several membrane proteins has been found to be correlated with
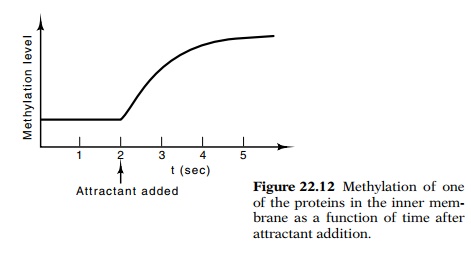
chemo-taxis behavior for some attractants. The addition of attractant to cells
leads to preferential methylation of one of a set of at least four mem-brane
proteins, products of genes named tsr,
tar, tap, and trg (Fig.
22.12). These proteins are the receptors and signal transducers for some of the
attractants and repellents to which E.
coli responds. For example the tar product is the receptor for the
attractants aspartate, maltose, and many repellents.
Signaling between the receptor proteins and the
intracellular chemo-taxis machinery requires sending a signal through the
membrane. The tar protein spans the
membrane. It has a domain of about one hundredof the N-terminal amino acids
outside the membrane, and about two hundred more amino acids inside the
membrane. The external portion binds the aspartate, but it is the internal
portion that is methylated by
methyl transfer from S-adenosyl methionine. As many
as five glutamic acid residues per receptor can be modified as the cells adapt
to the continued presence of attractant. The intracellular portion of the
pro-tein is highly homologous to other methyl-accepting chemotaxis pro-teins,
whereas the N-terminal portion is only weakly homologous to the other
chemotaxis receptors.
Increase in the concentration of an attractant
generates a conforma-tion change in one of these membrane-bound proteins. This
change is ultimately communicated to the flagellar motor. Increasing
methylation of the protein decreases its ability to send a signal for smooth
swimming or increases its ability to send a signal for tumbling. It is the
change in signal strength resulting from change in methylation that produces
adaptation.
For methylation to be part of the cell’s “memory,”
at least two enzymes are required: one is an enzyme to methylate and the other
is an enzyme to demethylate. More precisely, a methyltransferase and a
methylesterase must exist. Both of these have been found. They act on the
proper membrane proteins and are encoded by genes in the set of eight or nine
genes that are required for chemotaxis but are not required for motility.
Related Topics