Chapter: Genetics and Molecular Biology: Biological Assembly, Ribosomes and Lambda Phage
Determining Details of Local Ribosomal Structure
Determining Details of Local Ribosomal Structure
Consider the fundamental question of determining
which proteins are close neighbors in the ribosome. One direct approach to this
question is to crosslink two proteins on the intact ribosome with bifunctional
crosslinking reagents. If two ribosomal proteins are connected by the reagent
when they are in a ribosome, but not when they are free in solution, it can be
concluded that the proteins are near one another in the ribosome. This
technique is fraught with artifacts, however, and results from different
laboratories frequently do not agree, leading some to believe only those
crosslinking results that have been duplicated in more than two laboratories.
Many of the proteins that are crosslinked to each
other are proteins that depend on one another during assembly of the ribosomal
subunit. A few of these proteins are encoded in the same operons. In one case,
proteins that are adjacent to one another in the ribosome derive from adjacent
genes in the chromosome. For example, ribosomal proteins S4, S11, and S13 lie
in the same operon, S13-S11-S4. S4 and S13 and S13 and S11 crosslink, S4 and
S13 interact during assembly, and together they interact with S11 during
assembly.
The ability to reassemble ribosomes from their
isolated components greatly facilitates structural studies. A ribosome can be
partially assem-bled, for example, and then antibody against a component in the
immature ribosome can be added. If the presence of the antibody blocks the
subsequent association of a ribosomal protein added later, it is reasonable to
expect that the antibody directly blocks access of the protein to its site.
If all ribosomal proteins were spherical, their
complete spatial ar-rangement would be determined by knowing the distances
between the centers of proteins. Some of the requisite measurements can be made
with fluorescence techniques or slow neutron scattering. Fluorescent molecules
possess an absorption spectrum such that illumination by photons within this
wavelength band excites the molecule, which then emits a photon of longer
wavelength within what is called the emission spectrum of the molecule (Fig.
21.8).
In vitro assembly of ribosomes can be used to construct a
ribosomein which two of the proteins contain the fluorescent probes. By
illumi-nating the rebuilt ribosomes with light in the excitation spectrum of
the
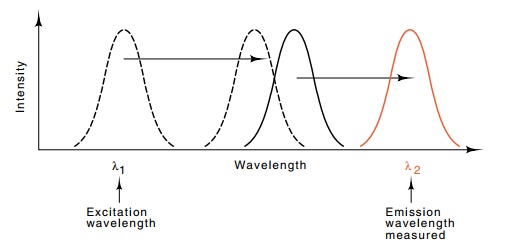
Figure
21.8 Spectra used in measuring
distances separating ribosomal pro-teins. Dotted line is the excitation and
emission spectrum of fluorescent molecule 1 and the solid line is the
excitation and emission spectra for molecule 2.
first molecule and measuring the strength of the
fluorescence in the wavelength of the emission spectrum of the second molecule,
the distance between the two fluorescent molecules can be determined. The
amount of light in the second emission spectrum varies as the sixth power of
the distance separating the molecules:
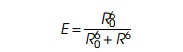
where R is the distance between the
fluorescent molecules and Ro
is a constant that depends on the orientations of the molecules, the spectral
overlap of the fluorescent emission and excitation spectra, and the index of
refraction of the medium separating the molecules. The method yields the most
reliable data for proteins separated by 25 to 75 Å; that is, the method is best
at determining the distances of nearest neighbors in the ribosome.
Neutron
diffraction is another method of measuring distances be-tween ribosomal
proteins. This method has yielded the most informa-tion and the most reliable
information on ribosome structure. It too relies on reassembly of ribosomal
subunits. Two proteins in the ribo-some are replaced by their deuterated
equivalents. These proteins are obtained from cells grown on deuterated medium.
Since the neutron scattering properties of hydrogen and deuterium are
different, an inter-ference pattern is generated by the presence in the
ribosome of the two proteins with different
scattering
properties. The angular separation in the peaks of the interference pattern can
be related to the distance separating the two altered proteins in the
reconstituted ribosome. Overall, the results of crosslinking, assembly
cooperativity, immune
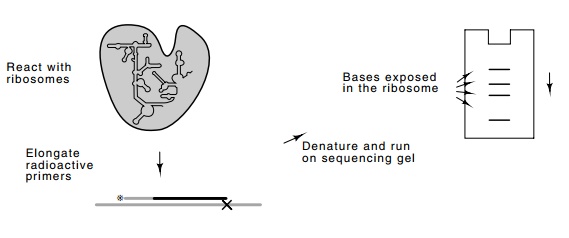
Figure
21.9 Technique for footprinting rRNA
in the intact ribosome. Theenhanced bands correspond to exposed bases. Normally
a control would be done reacting denatured rRNA and rRNA that is in an intact
ribosome. Then the bases protected and not protected are revealed by comparing
the band intensities from the free RNA and the RNA from the ribosomes.
Ribosomal RNA can also be footprinted like DNA.
Either bare RNA, RNA with a few proteins bound, or even an intact ribosome with
or without a bound protein synthesis inhibitor like streptomycin can be used.
The RNA is treated with chemicals like dimethylsulfate or kethoxal that react
with unprotected nucleotides. Then the RNA is purified and a DNA
oligonucleotide that will serve as a primer for reverse transcrip-tase is
hybridized. The elongation by reverse transcriptase ends at the modified bases,
and the locations of protected bases can be determined (Fig. 21.9).
Related Topics