Chapter: Biotechnology Applying the Genetic Revolution: Inherited Defects
Deleterious Tandem Repeats and Dynamic Mutations
DELETERIOUS TANDEM REPEATS AND DYNAMIC
MUTATIONS
Several genetic disorders are
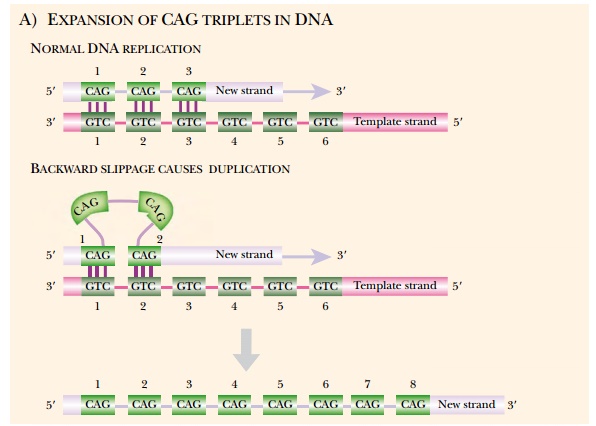
known where the defect is due to a tandem repeat of three bases within a
protein coding region. The three bases are usually CAG (in the coding strand),
which encodes glutamine. The wild-type alleles contain several CAG repeats that
give rise to a run of several glutamines within the encoded protein. The mutant
alleles have increased numbers of these repeats and give rise to proteins with
longer polyglutamine tracts (Fig. 16.4). Below a certain number (generally in
the range 5–30), these repeats are relatively harmless and stable. Beyond this
threshold, the mutations cause disease. In addition, the number of repeats is
unstable and tends to expand each successive generation, sometimes to over 100.
Hence these are sometimes referred to as dynamic
mutations. Generally, the higher the number of tandem repeats, the more
severe the pathogenic effects and the earlier the onset of disease.
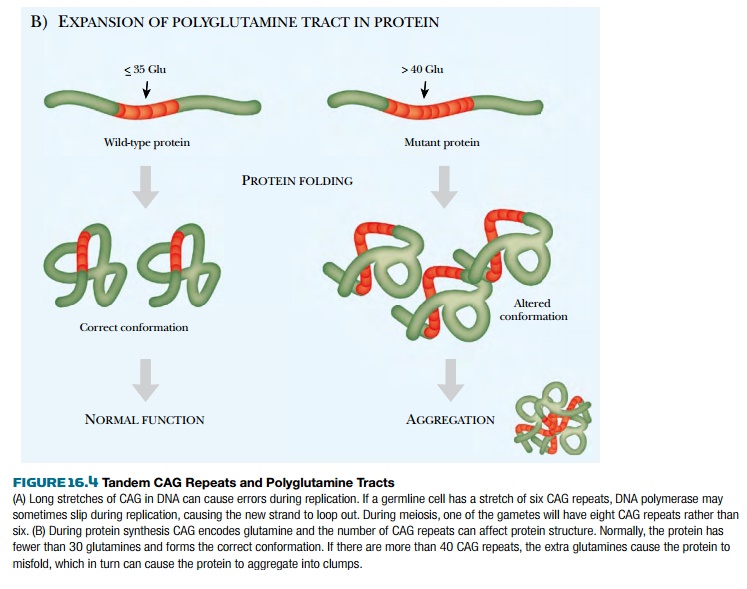
The known polyglutamine/CAG
repeat defects all cause neurodegenerative diseases with late onset. Almost all
are autosomal dominants. In all cases, the extended polyglutamine tracts cause
the proteins to clump into aggregates that ultimately kill nerve cells.
Different CAG repeat diseases affect different proteins that are expressed at
different levels in different nervous tissues. Consequently, the clinical
symptoms vary. Perhaps the best known is Huntington’s
disease, located on chromosome 4 at
4p16. Transport of vesicles in nerve cells appears to be affected and results in loss of control of the limbs, impaired
cognition, and dementia.
Unstable expanding tandem
repeats may also occur outside coding sequences. In some cases these are
harmless. In other cases they may affect the expression of nearby genes and
have devastating results. An example is fragile
X syndrome. The name refers to a fragile site within the long arm of the X
chromosome, whose effects can be seen under the microscope. The tandem repeats
may cause the chromosome to fragment into two parts or the two regions may
remain held together by a thin thread of material. Fragile X affects about 1 in
4500 males, and causes mental retardation. The syndrome is less common and much
less severe in females.
Although differences in
imprinting are also involved (see later discussion), this difference is largely
due to females having two X chromosomes, whereas males have only one.
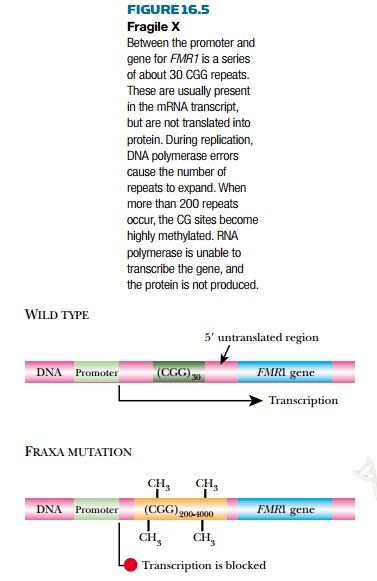
Fragile X syndrome is due to
tandem repeats of CGG located within the 5′-untranslated region of the FMR1
gene in band Xq27 on the affected X chromosome (Fig. 16.5). In the wild type
there are around 30 copies of CGG. As with the CAG repeats described earlier,
the number of repeats is unstable and tends to expand each successive
generation. If the number of repeats expands to 50 to 200, no symptoms are
observed, but such carriers often have children with more than 200 repeats, who
do show symptoms. The fully expanded version of the fragile X allele may have
up to 1300 CGG repeats. Although the CGG repeats are not translated into
protein, they act as CG islands and tend to be methylated. This abolishes
transcription of the FMR1 gene, which regulates the synthesis of proteins in
the synaptic junction of nerve cells. The defect causes mental retardation plus
other symptoms. Another fragile X site (fragile X site E or FRAXE) is located
in the FMR2 gene about 600 kb distal of the FMR1 gene (fragile X site A or
FRAXA).
Related Topics