Chapter: Genetics and Molecular Biology: Attenuation and the trp Operon
Attenuation and the trp Operon
The Aromatic Amino Acid Synthetic Pathway and
its Regulation
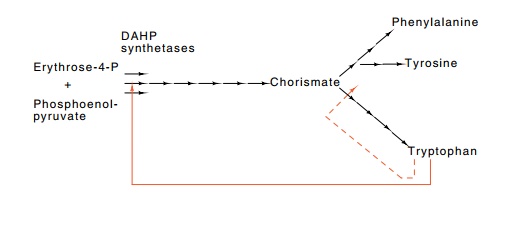
The first chemical reaction common to the synthesis
of tryptophan, tyrosine, and phenylalanine is the condensation of erythrose-4-P
and phosphoenolpyruvate to form 3-deoxy-D-arabino-heptulosonate-7-P, DAHP, a
reaction catalyzed by DAHP synthetase (Fig. 13.1). Due to its position at the
head of the aromatic amino acid biosynthetic pathway and to the fact that this
reaction is irreversible, it is logical that this reaction should be a key
point of regulation. Indeed, this expectation is met. Escherichia coli regulates both the amount and activity per
mole-cule of DAHP synthetase as a function of the intracellular levels of the
aromatic amino acids.
Figure
13.1 Outline of the aromatic amino
acid biosynthetic pathway inEscherichia
coli. Each arrow represents an enzymatic step. Tryptophan feed-back
inhibits one of the DAHP synthetases, AroH, as well as the first enzyme of the
pathway committed to tryptophan synthesis.
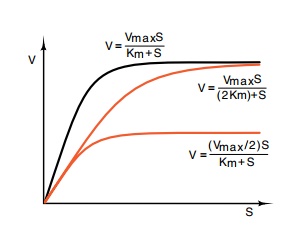
Figure
13.2 Two possibilities for feed-back
inhibition of an enzyme. The top curve is enzyme velocity as a function of
substrate concentration, and the equa-tion is the activity of an uninhibited
enzyme as a function of its Km, Vmax, and substrate concentration S.
A double regulation of total DAHP synthetase activity is logical. While regulation of the enzyme level minimizes unnecessary consumption of amino acids and energy in the synthesis of the
enzyme, this type of regulation is incapable of producing appreciable changes
in the enzyme levels or of total enzyme activity on time scales shorter than
minutes. A much more rapidly-responding regulation mechanism is also necessary
to adjust the synthesis rates of tryptophan, tyrosine, and phenylalanine on time
scales of seconds.
In addition to stabilizing the activity of the
synthetic pathway against random fluctuations, a rapidly responding regulation
would fine-tune the biosynthetic flow rates of the aromatic amino acids and
would be able to respond rapidly to growth rate changes generated by changes in
the nutrient medium. Feedback inhibition of an enzyme’s activity meets the
requirements, as this mechanism can alter an enzyme’s activity in milliseconds.
Feedback inhibition is an example of an allosteric
interaction in which accumulation of the product of the pathway leads to
inhibition of the activity of an enzyme in that pathway. This is a specific
example of an allosteric interaction in which a molecule dissimilar in shape to
the substrates of an enzyme can bind to the enzyme, usually at a site on the
enzyme far from the active site, and can generate conformational changes that
alter the catalytic activity of the enzyme. Feedback inhibi-tion can reduce an
enzyme’s activity in either of two fundamental ways (Fig. 13.2). The
tryptophan-sensitive DAHP synthetase is feedback-in-hibited largely as a result
of a change in its Vmax, whereas the first enzyme of the pathway used
solely for tryptophan synthesis, anthra-nilate synthetase, is an example of the
other possibility. It is feedback-inhibited by tryptophan via a change in its Km.
Bacillus
subtilus possesses a single DAHP
synthetase whose synthesisand activity is regulated by the three aromatic amino
acids. Escherichiacoli, however,
possesses three different DAHP synthetases. The activityof one, the AroH
protein, is feedback-inhibited by tryptophan, another is feedback-inhibited by
tyrosine, and the third is feedback-inhibited by phenylalanine. Only if the
cell’s growth medium possesses all three amino acids is all DAHP synthetase
activity within an E. coli cell fully
inhibited
This is an example of the fact that different
microorganisms possess different overall schemes for the regulation of
tryptophan synthesis. On one hand, it is possible that the different evolutionary
niches occupied by different microorganisms require these different schemes. On
the other hand, perhaps one scheme is no better than another, and the different
ones just happened to have evolved that way. Either way, this diversity means
that the overall scheme for regulation of tryptophan synthesis in E. coli
is not the only one that works.
Related Topics