Chapter: Biotechnology Applying the Genetic Revolution: Basics of biotechnology
Viruses Used in Genetics Research
VIRUSES
USED IN GENETICS RESEARCH
Viruses are entities that
border on living. But unlike genuine living organisms, viruses cannot survive
outside a host organism. Viruses are pathogens that invade host cells and
subvert them to manufacture more viruses. Viruses are simple in principle and
yet very powerful. They consist of a protein shell called a capsid surrounding a genome made of RNA
or DNA. The particle is called a virion
and, unlike a living cell, has no way to make its own energy or duplicate its
own genome. The virus relies on the host to do this work.
Viruses come in many
different types and can inhabit every living thing from bacteria to humans to
plants. Viral diseases in humans are extremely common, and most cause only
minor symptoms. For example, when rhinovirus invades, the victim ends up with a
runny nose and other cold symptoms and usually feels miserable for a few days.
However, viruses do cause a significant number of serious diseases, such as
AIDS, smallpox, hepatitis, or the recently identified Ebola.
When viruses invade bacteria,
the infected bacteria usually die. Bacterial viruses are called bacteriophage or phage, and they normally destroy the bacterial cell in the process
of making new viral particles. When bacteria grow on an agar plate, they form a
hazy or cloudy layer (a bacterial lawn)
over the top of the agar. If the culture of bacteria is infected with
bacteriophage, the viruses eat holes or plaques
into the bacterial lawn, leaving clear zones where all the bacteria were
killed.
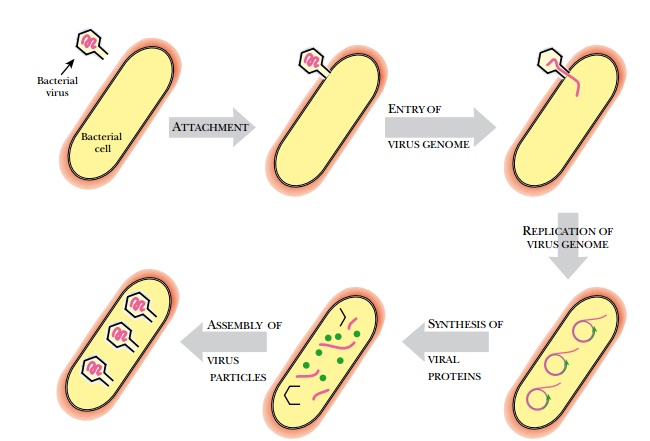
Bacteriophages, like other
types of viruses, have the following stages of their life cycle (Fig. 1.23):
(a)
Attachment of the virion to the correct host cell
(b)
Entry of the virus genome
(c)
Replication of the virus genome
(d)
Manufacture of new virus proteins
(e)
Assembly of new virus particles
(f)
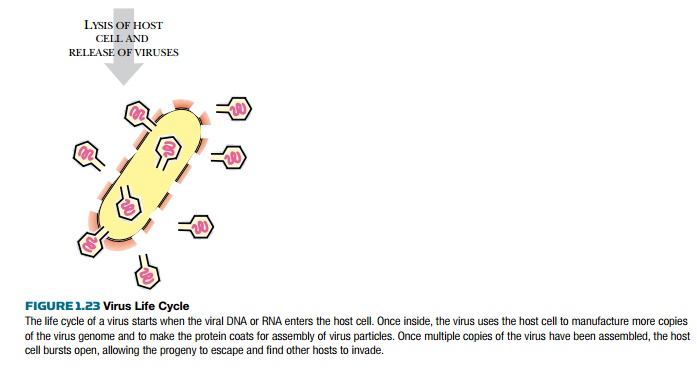
Release of new virions from the host
Not every virus kills the
host cell, and in fact, many viruses have a latent phase where they lie dormant
within the cell, not producing any proteins or new viruses. Latency, as it is called in animal
cells, is also called lysogeny when
referring to bacteria. (In contrast, the phase of viral growth where the host
cell is destroyed is called the lytic
phase.) Sometimes a virus becomes latent by inserting its genome into the
genome of the cell. The viral genome integrates into a host chromosome and
remains inactive until some stimulus triggers it to reactivate. The integrated
virus is called a provirus (or a prophage if the virus invades
bacteria).
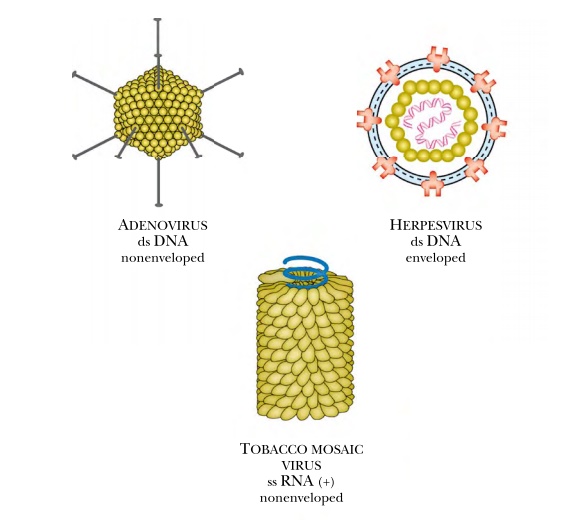
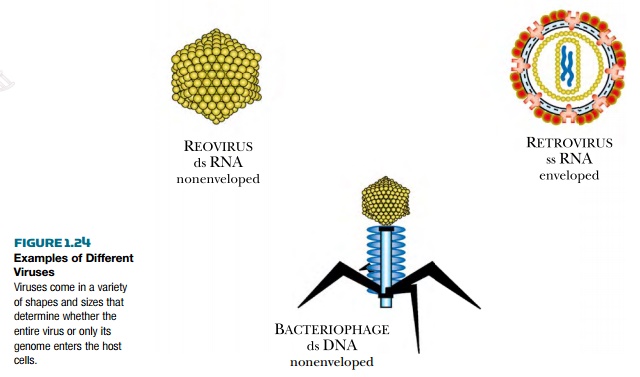
The great variety of viruses
can be divided into groups based on capsid shape or the type of genome. The
three major shapes are spherical, filamentous, and complex. Spherical viruses
actually have 20 flat triangular sides and are thus icosahedrons. Complex
viruses come in various shapes, but some have legs that attach to the host
cell, a linear segment that injects the DNA or RNA genome into the host, and a structure
that stores the viral genome. This type of complex virus is common among
bacteriophages, several of which are widely used in molecular biology research.
Bacteriophage T4, lambda, P1, and Mu all look like the Apollo lunar landers
(Fig. 1.24).
Viral genomes are varied in
size, but all contain sufficient genetic information to get the host cell to
make more copies of the virus genome and make more capsid proteins to package
it. At the very least, a virus needs a gene to replicate its genome, a gene for
capsid protein, and a gene to release new viruses from the host cell.
Bacteriophage Qβ infects bacteria; its entire
genome is only 3500 base pairs, and the entire genome consists of only four
genes. On the other hand, large complex viruses may have more than 200 genes
that are used at different
times after infecting the
host cell. The genes are then divided into categories based on when they are
active. Some genes are considered early
genes and are active immediately after infecting the host, whereas late genes are active only after the
virus has been inside the host cell for some time.
Viral genomes are either made
from DNA or RNA, can be double-stranded or single-stranded, and can be circular
or linear. When viruses use a single strand of RNA as genome,
this can either be the positive or plus (+) strand or the negative
or minus (–) strand. The positive
strand corresponds to the coding strand and the negative strand to the template
strand. When a positive-strand RNA virus injects its genome into the host, the
RNA can be used directly as a messenger RNA to make protein. If the RNA virus
has a negative-strand genome, the RNA must first be converted into
double-stranded form (the replicative
form or RF). Then each strand is
used: the negative strand is used as a template to make more positive-stranded genomes, and the positive strand is
used to make proteins.
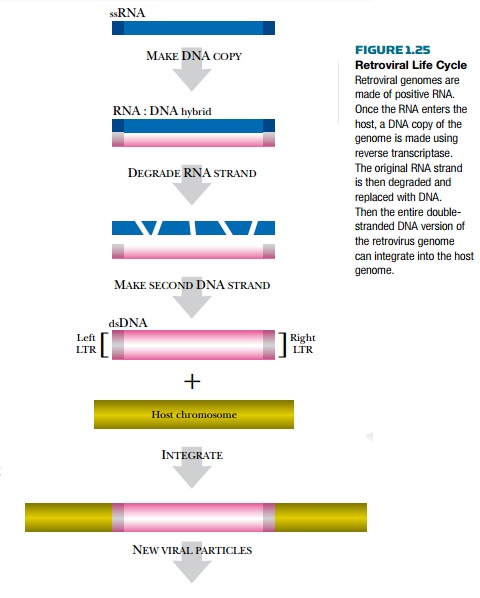
Some viruses actually use both RNA and DNA versions of their genome (Fig. 1.25). Retroviruses infect animals and include such members as HIV (human immunodeficiency virus). The genome inside a retrovirus particle is a single-stranded RNA that is converted to DNA once it enters the host. Reverse transcriptase is the enzyme that manufactures the DNA copy of the RNA genome and is used extensively in molecular biology and genetic engineering. Once the DNA copy is made, it is inserted into the host DNA using two repeated DNA sequences at the ends called long terminal repeats (LTRs). Once integrated, the retrovirus becomes part of the host’s genome. This is why there is no complete cure for acquired immunodeficiency syndrome (AIDS); the host can never rid itself of the retroviral DNA once it becomes integrated. The viral genes then direct the synthesis of new viral particles that infect neighboring cells. Reverse transcriptase is an example of a viral gene product that is synthesized by the host and packaged inside the virions for use in the next infection cycle.
Retroviral genomes have three major genes, gag, pol, and env, as well as several minor genes. The tat and rev genes regulate the expression of the other retroviral genes. Nef, vif, vpr, and vpu encode four accessory proteins that block the host cell’s immune defense and increase the efficiency of virus production. Gag, pol, and env each give single mRNA transcripts that encode multiple proteins. Gag encodes three proteins involved with making the capsid. Pol gives three proteins, a protease that digests other proteins during particle assembly, reverse transcriptase that makes the DNA copy of the genome, and an integrase that integrates the viral DNA into the host chromosome. Env codes for two structural proteins; one forms the outer spikes and the other helps the virus enter the host cell.
Related Topics