Chapter: Microbiology and Immunology: Virology, Virus: General Properties of Viruses
Nomenclature and Taxonomy of Viruses
Nomenclature and Taxonomy of Viruses
The initial classification of viruses was haphazard and was based on the symptomatology of the disease they caused or on the sites of their isolation. They were classified as (a) pneumo-tropic, those affecting the respiratory tract (e.g., influenza and common cold); (b) neurotrophic, affecting the nervous system (e.g., poliomyelitis and rabies); (c) dermatotropic, affecting the skin (e.g., smallpox and chickenpox); and (d) viscerotropic, affecting visceral organs (e.g., hepatitis, yellow fever). This system of classification was convenient but not scientifically acceptable, because the same virus may cause more than one disease depending on the tissue affected.
Since 1966, a systematic classification of viral taxonomy and nomenclature was carried out with the formation of International Committee on the Taxonomy of Viruses (ICTV). Since then, ICTV has been introducing systematic approach for classification and nomenclature of the viruses. The ICTV has been grouping viruses into families based on (a) type of nucleic acid they possess, (b) means of replication, and (c) morphology (e.g., presence or absence of membrane envelope).
The suffix virus is used for genus names, viridae for family names, and ales for order names. In formal usage, the family and genus names are used in the following manner: for example, fam-ily Rhabdoviridae, genus Lyssavirus, human rabies virus. A viral species is a group of virus that shares the same genetic informa-tion and ecological niche. These viral species are designated by descriptive common names, such as human herpesvirus, with subspecies, if any, designated by a number (e.g., HHV-1).
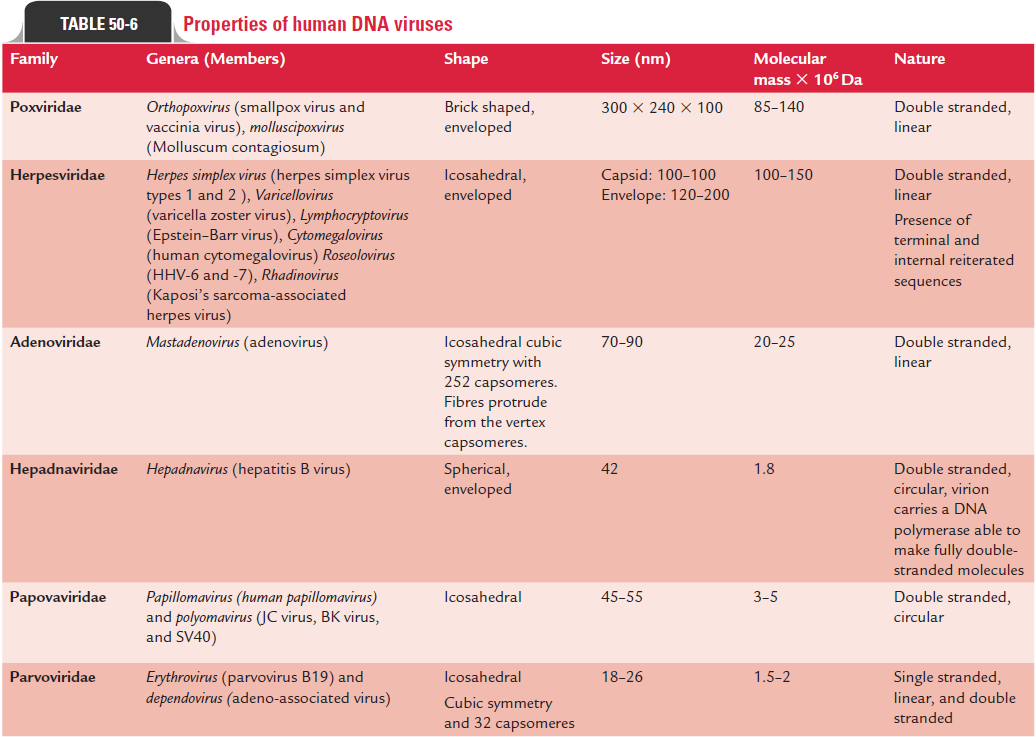
Depending on the type of nucleic acids viruses possess, they are classified into two groups: deoxyriboviruses, which contain DNA (DNA virus) and riboviruses, which contain RNA (RNA virus). DNA viruses associated with human diseases are divided into six families (Table 50-6). The RNA viruses associated with human diseases are classified into at least 13 families (Table 50-7).
DNA Viruses
These belong to the following families (Table 50-6):
Adenoviridae: The members of the family Adenoviridaeare medium-sized viruses measuring 20–90 nm in size. These viruses are nonenveloped, icosahedral viruses with 252 cap-someres. They are so named because they were first isolated from adenoids. These viruses are mostly associated with acute respiratory diseases.
Poxviridae: These are large-sized, brick-shaped viruses mea-suring 300 3 240 3 100 m in size. They have a complex structure with a core containing a single linear molecule of double-stranded DNA genome. The pox (pox: pus-filled lesions) viruses are associated with skin lesions. The viral components are syn-thesized and assembled in the cytoplasm of infected host cells.
Herpesviridae: These are medium-sized icosahedral nucleo-capsid viruses (100 nm) containing 162 capsomeres. They are enveloped viruses containing linear, double-stranded DNA. They are named after spreading (herpetic) appearance of cold sores. The viruses multiply in the nucleus of the host cells.
Papovaviridae: These are small (40–55 nm) viruses con-taining double-stranded DNA with 72 capsomeres. They are nonenveloped viruses. They replicate in the nucleus of host cell along with host cell chromosome. This may cause host cells to proliferate, resulting in a tumor. Papovaviruses are acronyms to papillomas (warts), polyomas (tumors), and vac-uolation (cytoplasmic vacuolation produced by some of these viruses).
Hepadnaviridae: Hepadnaviridae (hepa: liver;dna: DNA core)are so named because they cause hepatitis and contain DNA as genome. These viruses differ from other DNA viruses by synthe-sizing their DNA by copying RNA using reverse transcriptase. Human hepatitis B virus, an important virus associated with human disease, is included in this family.
RNA Viruses
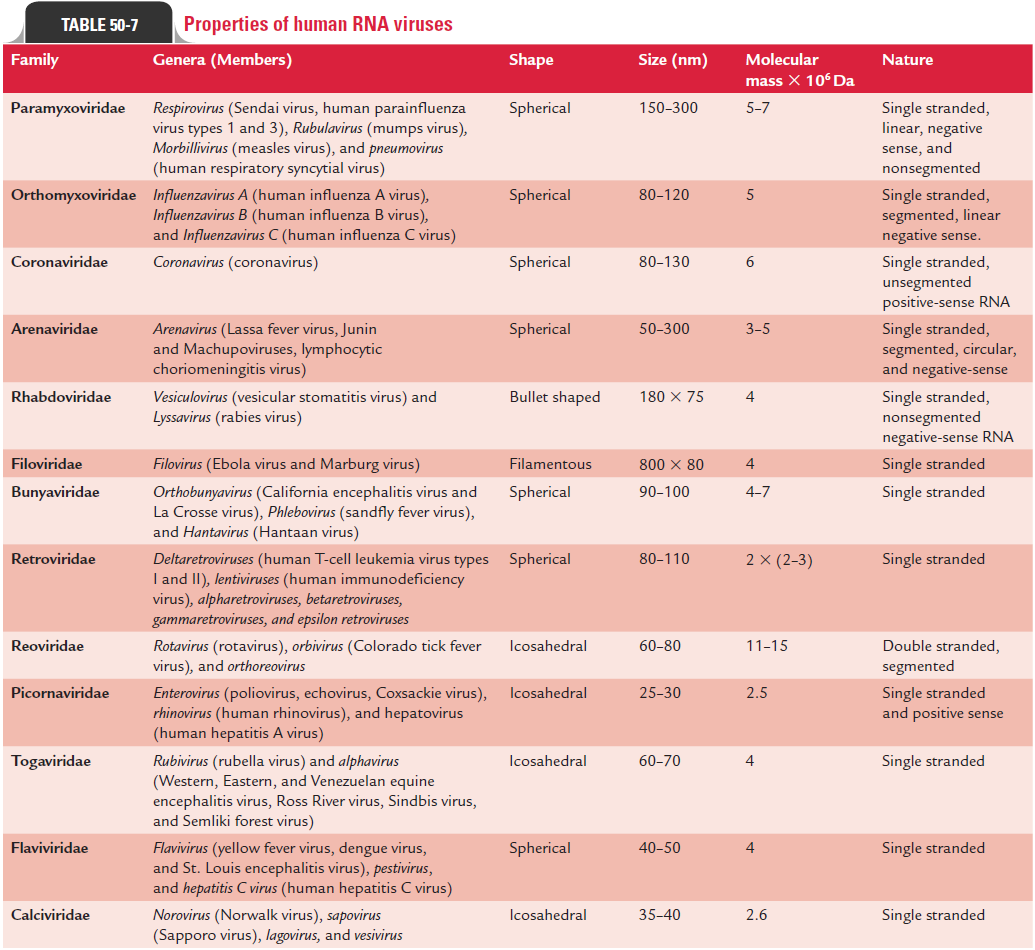
These belong to the following families (Table 50-7):
Togaviridae: These viruses include arboviruses and alphavi-ruses. Most of these viruses multiply in arthropods as well as in vertebrates. Togaviridae (toga: envelope) are enveloped viruses containing single-stranded RNA genome. These viruses are small spherical viruses measuring 40–70 nm in size.
Rhabdoviridae: Rhabdoviruses (rhabdo: rod) are bullet-shapedviruses. They are enveloped, measure 130–300 3 20 nm in size, and contain a single-stranded RNA.
Reoviridae: They are icosahedral, nonenveloped viruses mea-suring 60–80 nm in size. They contain double-layered capsid enclosing 10–12 segments of double-stranded RNA. Their name is derived from the first letters of respiratory, enteric, and orphan. When first discovered, the viruses were not associ-ated with any diseases, hence were called orphan viruses. These viruses are now known to cause respiratory and intestinal infections.
Retroviridae: Retroviruses (re: reverse,tr: transcrip-tase) viruses are so named because characteristically they possess the enzyme, reverse transcriptase RNA-dependent DNA polymerase. They are icosahedral, enveloped viruses measuring 100 nm in size. Many of these viruses are asso-ciated with tumors in infected hosts. One of the genera, Lentivirus includes the subspecies HIV-1 and HIV-2, the causative agents of aquired immunideficiency syndrome (AIDS).
Picornaviridae: Picornaviruses (pico: small) are the small-est viruses, measuring 20–30 nm in size. They are nonenvel-oped, icosahedral viruses with single-stranded RNA genome. These include three genera (Enterovirus, Rhinovirus, and Hepatovirus) of medical importance.
Orthomyxoviridae: These are medium-sized (80–120 nm)viruses. They are spherical and elongated, enveloped viruses consisting of single-stranded but segmented (eight segments) RNA genome. Influenza virus is the only virus of medical importance belonging to this group.
Paramyxoviridae: These are pleomorphic, enveloped virusesmeasuring 150 nm in size. They contain nonsegmented, single-stranded, linear RNA. Three genera have been described: Paramyxovirus, Morbillivirus, and Pneumovirus.
Bunyaviridae: These are enveloped, spherical viruses measur-ing 90–100 nm in size. The genera of medical importance include Bunyavirus, Hantavirus, Uukuvirus, Phlebovirus, and Nairovirus.
Arenaviridae: They are spherical, pleomorphic viruses withvariable sizes (50–300 nm). They contain electron-dense, chro-mosome-like particles giving a sandy appearance; hence they are named arenaviruses (arena: sand).
Calciviridae: These are naked nonenveloped viruses. They aresmall and spherical, and measure 35–39 nm in size. They show 32 cup-shaped depressions arranged in symmetry.
Filoviridae: They are long filamentous, enveloped viruses withvariable sizes. They contain single-stranded RNA genome. The Marburg and Ebola virus are the viruses of medical importance.
Prions
Prions are infectious particles, which can transmit a disease. These prions are composed chiefly a protein without any detect-able nucleic acid. This is a new name widely accepted for the old name slow viruses.
The term prion was coined by Prusiner for proteinaceous infectious particles. Unlike conventional viruses, the prions apparently have no virion structure or genomes and evoke no immune response in the infected host. These are extremely resistant to inactivation by heat, disinfectants, and radiation.
The prions are causative agents of slow viral infections, such as subacute spongiform encephalopathy. After long incubation period of years, they produce a progressive dis-ease that causes damage to the central nervous system, lead-ing to subacute spongiform encephalopathy.
Viroids
Viroids are protein-free fragments of single-stranded, circular RNA that cause disease in plants. The term viroids was coined by Triener (1971) to describe a new subclass of subviral agents having a genome very much similar to those of known viruses; they consist of very short pieces of naked RNA: 300–400-nucle-otides long with no protein coat. The RNA does not code for any proteins. The low-molecular-weight RNA is resistant to heat and organic solvents but sensitive to nucleases. So far, viroids have been identified as pathogens of only plants; these were first identified in the potato spindle tuber disease. The viroids are yet to be linked to any disease in humans.
Related Topics