Chapter: Plant Biology : Interactions between plants and other organisms
Nitrogen fixation in Plant Biology
NITROGEN FIXATION
Key Notes
Nitrogen fixation
Nitrogen gas cannot be used directly by plants but is fixed to nitrogencontaining compounds by free-living or symbiotic bacteria and cyanobacteria. Nitrogen-fixing legumes form one of the world’s largest families, dominating many plant communities.
The infection process
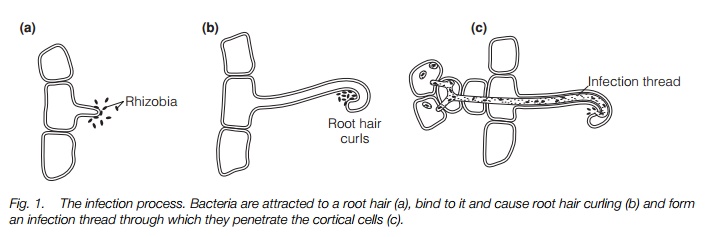
The legume root hair secretes a chemo-attractant which causes the bacteria to accumulate. They cause root-hair curvature and enter the root cortex by an infection thread. Cell division is stimulated to form a nodule with vascular connections to the plant.
Molecular biology of nitrogen fixation
The complex interaction of host and bacterium requires the coordinated action of NOD genes in the host that encode nodulation and leghemoglobin, and nod, nif and fix genes in the bacterium that encode infection, host specificity and components of nitrogen-fixation.
Biochemistry of nitrogen fixation
Nitrogen fixation requires 16 moles of ATP per mole of nitrogen and almost anaerobic conditions created by the oxygen binding protein leghemoglobin. The bacteria in the cytoplasm are surrounded by the peribacteroid membrane. Nitrogen-fixation is catalyzed by dinitrogenase in three stages: (i) reduction of the Fe-protein; (ii) reduction of the MoFe protein by the Fe protein (requires ATP); (iii) reduction of nitrogen by the MoFe protein. Nitrogen is exported in high-nitrogen containing compounds such as amino acids or ureides.
Nitrogen fixation
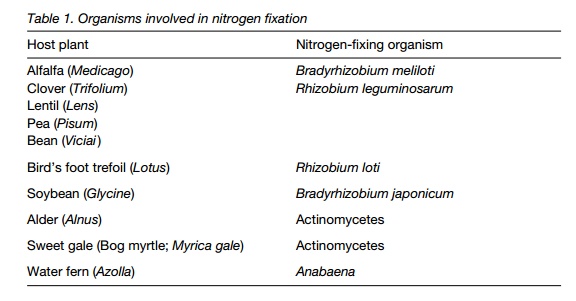
Nitrogen (N2) is abundant in the aerial and soil environment, but unlike oxygen cannot be used directly . Nitrogen-fixation occurs in some free-living microorganisms (for instance some cyanobacteria, bacteria and archaebacteria), but the most complex systems are seen in symbiotic association with some species of plants (Table 1). Nitrogen-fixing symbioses with the nitrogen-fixing bacteria Rhizobium and Bradyrhizobium are important in agricultural crops (peas, beans and other legumes, and clover in pasture land). In the Far East, the floating water fern Azolla that forms a nitrogen-fixing symbiosis with Anabaena, a cyanobacterium, is used to fertilize rice-paddy cultivation. Alder (Alnus) trees, sweet gale (Myrica gale) and mountain lilacs (Ceanothus) form nitrogen-fixing symbioses with actinomycetes (a different group of bacteria).
The legume–rhizobium symbiosis has been studied in most detail and will be considered here. Legumes (members of the family Fabaceae) are the most widespread and important plants with nitrogen-fixing root nodules and form one of the world’s most abundant plant families, including many trees, such as Acaciaspecies. The ability of many legumes to fix atmospheric nitrogen has been one of the major factors in their success. Many plant communities, particularly those on soils that are poor in nitrogen, are dominated by legumes, and they can enhance the nitrogen status of the soil for other plants.

The infection process
The legume root hair secretes a chemo-attractant which cause the bacteria to accumulate (Fig. 1). The bacteria secrete lipochito-oligosaccharides (NODfactors, see below) that cause more root hairs to be formed and alter root metabolism. They cause root-hair curvature and the bacteria then attach to the hair by sugar-binding proteins called lectins. An infection thread is then formed, by which the bacteria pass through the root hair to the root cortex, where they proliferate. Cell division is stimulated to form a nodule within which nitrogen fixation occurs. The nodule has good vascular connections through which carbohydrates are supplied to the nodule and nitrogen-containing compounds are exported to the plant.
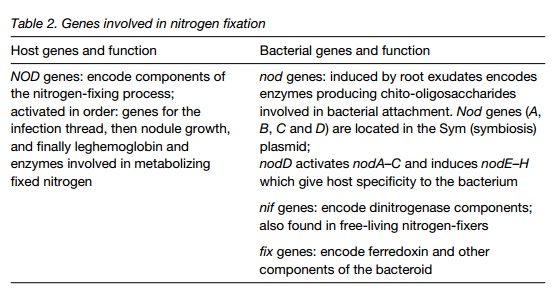
Molecular biology of nitrogen fixation
The complex interaction of host and bacterium requires the coordinated action of a number of key genes in both organisms. Table 2 lists these genes and their functions. The NOD gene family in the host encodes aspects of nodule formation and infection and the production of leghemoglobin, which binds oxygen, lowering the concentration of O2 in the nodule. The nod family in the bacterium encodes enzymes that synthesize NOD factors (see above). One nod gene, nod is activated by root exudates and produces a product that regulates the other nod genes. Later in infection, the nif and fixgenes are active in the rhizobium. These produce the enzymes and electron transport pathway required for nitrogen-fixation.

Biochemistry of nitrogen fixation
Nitrogen fixation is a very energy demanding process, requiring at least 16 moles of adenosine triphosphate (ATP) for each mole of reduced nitrogen. It also requires almost anaerobic conditions as the key enzyme complex, dinitrogenase, is rapidly inactivated by oxygen. In the nitrogen-fixing nodule, the energy required is supplied by the plant, and low oxygen conditions by the oxygen binding protein leghemoglobin. The bacteria are contained in the cytoplasm of cells in the nodule as bacteroids surrounded by a double membrane, the peribacteroid membrane. The bacterium supplies the plant with reduced nitrogen compounds such as ammonium and amino acids. The plant benefits by increased growth and the bacterium from a food supply and enhanced environment in which to grow and replicate. The overall equation of nitrogen fixation is:
8H+ + 8e–+N2 + 16ATP ®2NH3 + H2 + 16ADP + 16PI
The process is catalyzed by the enzyme dinitrogenase, a protein complex made of two components: a large component, the MoFe protein and a smaller component, the Fe protein. Dinitrogenase functions in stages:
(i) reduction of the Fe-protein (usually by the electron donor ferredoxin);
(ii) reduction of the MoFe protein by the Fe protein (this step requires ATP);
(iii) The MoFe protein then reduces nitrogen (N2 + 8H+ ®2NH3 + H2).
After fixation, nitrogen is exported to the plant in the xylem flow, not in the form of ammonium, but as high-nitrogen containing compounds such as amino acids or ureides, depending on the species.
Related Topics