Chapter: Microbiology and Immunology: Virology, Virus: Orthomyxoviruses
Influenza Viruses: Properties of the Virus
Influenza Viruses
Influenza viruses are classic respiratory viruses. They cause influenza, an acute respiratory disease, with well-defined systemic symptoms. Influenza is an acute infectious disease of the respiratory tract that occurs in sporadic, epidemic, and pandemic forms.
Properties of the Virus
◗ Morphology
Influenza viruses show following features:
· Influenza viruses are spherical or filamentous, enveloped particles 80–120 nm in diameter.
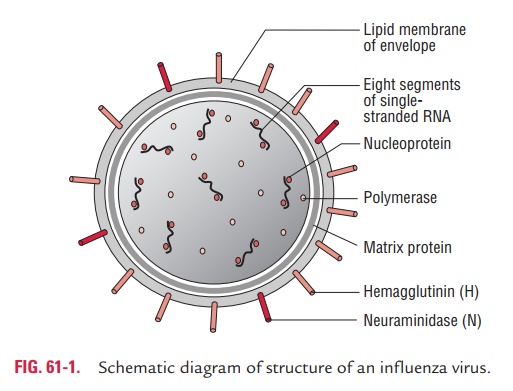
· Influenza virus is composed of a characteristic segmented single-stranded RNA genome, a nucleocapsid, and an envelope (Fig. 61-1).
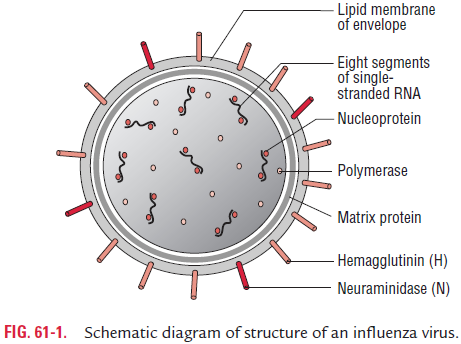
· The viral genome is a single-stranded antisense RNA. The genome consists of an RNA-dependent RNA polymerase, which transcribes the negative-polarity genome into mRNA. The genome, therefore, is not infectious. The viral RNA has a molecular weight of 5 million daltons and a length of 13,600 nucleotides. Characteristically, it is segmented and consists of seven or eight segments. These segments code for different proteins which are NS1, NS2, NP, M1, M2, M3, HA, and NA.
· The genome is present in a helically symmetric nucleocapsid surrounded by a lipid envelope. The envelope has an inner membrane protein layer and an outer lipid layer. The mem-brane proteins are known as matrix or M protein and are composed of two components M1 and M2.
· Two types of spikes or peplomers project from the envelope: (a) the triangular hemagglutinin (HA) peplomers and (b) the mushroom-shaped neuraminidase (NA) peplomers.
◗ Viral replication
Influenza virus, hepatitis delta virus, and retroviruses are the only RNA viruses that have an important stage of their replica-tion cycle in the nucleus. Infection of the host cell begins by adsorption of the cell by influenza virus, which is mediated through the HA. HA is first cleaved by an extracellular protease to a modified HA that actually mediates the attachment of the virus to the cell surface. Once inside the cell, the virus uncoats within the endosome.
In the nucleus of the host cell, the virion RNA polymerase transcribes the eight-genome segments into eight viral mRNAs. Most of the viral mRNAs, however, move out of the nucleus into the cytoplasm, where they are translated into viral proteins. Some of the viral mRNAs continue to remain in the nucleus and serve as the templates for synthesis of the negative-strand RNA genomes for the progeny virions. In the nucleus also, two proteins, namely, nucleoprotein (NP) and matrix protein are synthesized, which then bind with the RNA genome of the viral progeny and form a complex, which is subsequently trans-ported to the cytoplasm.
Matrix protein mediates the interaction of the nucleocapsid with the envelope, and finally, the virion is released from the cell by budding from the cell membrane at the site where the HA and NAs are present.
◗ Antigenic and genomic properties
Influenza viruses have two types of antigens:
Group-specific antigens: The ribonucleoprotein (RNP)antigen, or the “soluble” antigen, or the internal antigen is the group-specific antigen. Influenza viruses are divided into types A, B, and C on the basis of variation in this nucleoprotein antigen.
Type-specific antigens: The surface antigen, or “viral” anti-gen, or “V antigen” is composed of two virus-encoded proteins, HA and NA, which are the type-specific antigens (Fig. 61-2).
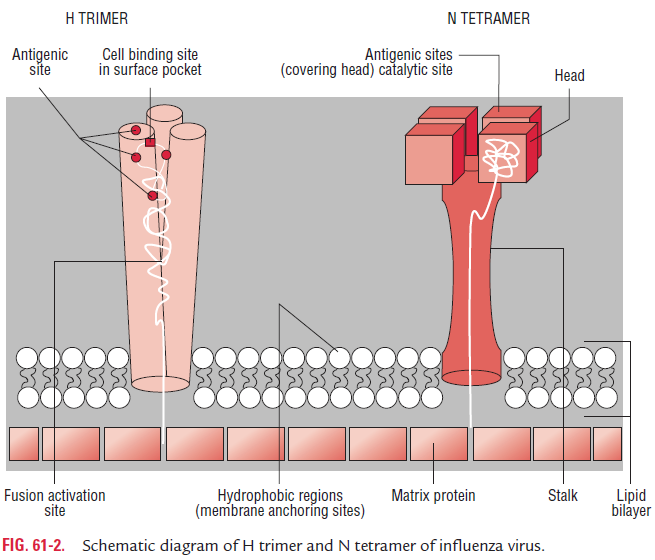
Hemagglutinin:HA is a trimer and is composed of two poly-peptides, HA1 and HA2, responsible for hemadsorption and hemagglutination. The hemagglutinin consists of 500 spikes, each measuring 12 nm in length. The triangular-shaped HA is inserted into the virus membrane by its tail end. The distal end, which contains five antigenic sites (designated as HA1–HA5), is responsible for binding of virion to host cells.
Influenza viruses adsorb many avian and mammalian eryth-rocytes. Hemagglutinin binds with the neuraminic acid (sialic acid) cell receptor, the cell surface glycoprotein receptor, and initiates the infection in the host cell. Adsorption of erythro-cytes occurs at 4°C, but at 37°C there is detachment of the red cells due to destruction of the glycoprotein receptors by the viral enzyme, neuraminidase. The hemagglutinin agglutinates certain red blood cells, which is inhibited by the neutralizing antibodies. This forms the basis of the hemagglutination inhi-bition test used in the serodiagnosis of influenza.
Hemagglutinin has potency to undergo antigenic variations. The nucleotide and amino acid sequences of the polypeptides, HA1 and HA2, undergo radical changes in antigenic shift. In antigenic drift, only minor changes take place in the composi-tions of HA antigenic sites.
Neuraminidase: The NA is a glycoprotein and tetramer. Itconsists of 100 mushroom-shaped spikes.The mushroom-shaped NA is inserted into the virus membrane by its hydro-phobic tail end. The distal end contains antigenic as well as enzymatically active sites. The NA causes hydrolysis of N-acetyl neuraminic acid or sialic acid residues present on theglycoprotein receptors on red cells, hence causes elution or detachment of the cells adsorbed to virion particles. The func-tion of the neuraminidase is to cleave the neuraminic acid and to release progeny virions from the infected host cells. The neuraminidase also degrades the mucus layer, thereby exposing the epithelial membrane of the respiratory tract for infection by the virus.
All the strains of influenza A, B, and C share the common internal proteins, such as nucleoproteins and membrane proteins. They, however, differ in their surface proteins, such as HAs and NAs. The influenza viruses are further subdivided into subtypes, A0, A1, A2, and A3, and these subtypes into strains.
◗ Antigenic variations
Antigenic variation is a unique feature of influenza virus. The surface antigens HA and NA show variations and are primarily responsible for antigenic variations exhibited by influenza viruses. The internal RNP antigen and M protein are stable, hence do not contribute to the antigenic varia-tions. Antigenic variations are of two types: antigenic shift and antigenic drift.
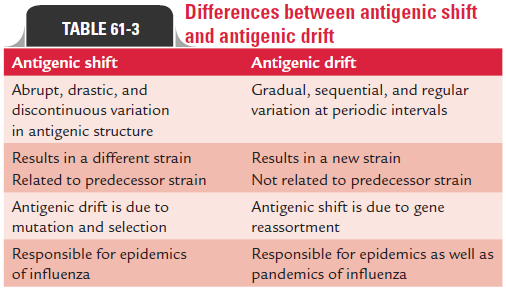
Antigenic shift: The abrupt, drastic, discontinuous change iscalled the antigenic shift. This occurs due to major antigenic changes in HA or NA antigens, and is caused by replacement of the gene for HA by one coding for a completely different amino acid sequence. The antigenic shift is characterized by alteration of virtually all the antigenic sites of the HA. This occurs inde-pendently in the HA and NA.
Antigenic drift: The gradual, sequential, regular antigenicchange in influenza virus is known as antigenic drift. This occurs due to minor antigenic changes in the HA or NA occurring at frequent intervals. This is caused even by a single mutation affecting HA glycoprotein. The antigenic drift is characterized by changes in certain epitopes in the HA, while others are being conserved.
Differences between antigenic shift and antigenic drift are summarized in Table 61-3.
◗ Gene reassortment
Because the influenza virus genome is segmented, genetic reassortment can occur when a host cell is infected simultane-ously with viruses of two different parent strains. This process of genetic reassortment accounts for the periodic appearance of the novel types of influenza A strains that cause influenza pandemics.
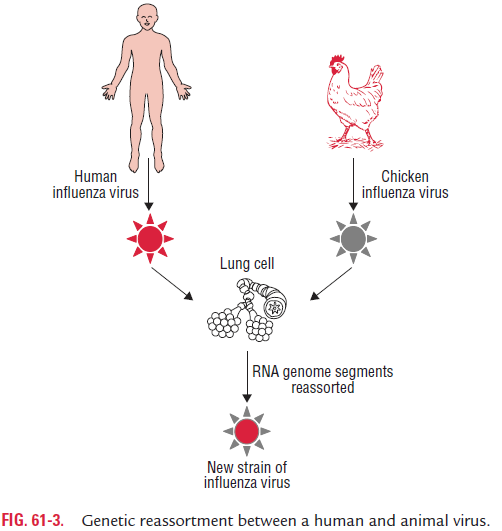
Influenza viruses of animals, such as aquatic birds, chickens, swine, and horses show high host specificity. These animal viruses are the source of the RNA segments that encode the antigenic shift variants that cause epidemics among humans. For example, if a person is infected simultaneously by an avian and human influenza strains, then it is possible that a genetic reassortment could occur in infected cells in humans. The reassortment could lead to emergence of a new human influenza A virus, the progeny of which will contain a mixture of genome segments from the two strains (e.g., a new variant of human influenza A virus bearing the avian virus HA) (Fig. 61-3).
Many studies have conclusively demonstrated that the aquatic birds (such as water fowl) are a common source of these new genes. The pigs act as mixing vessels, where these virulent genes of water fowl mix with the genome of influenza virus giving rise to new variant of influenza virus.
◗ Designation of influenza viruses
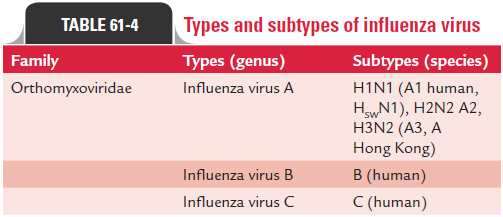
Influenza virus type A can be classified into subtypes based on the variations in their surface antigens (Table 61-4). The WHO proposed a new system of classification in 1971 and was later modified, which takes into account the nature of both the surface antigens. According to this, the complete designation of a strain will include the (a) type, (b) place of origin, (c) serial number, and (d) year of isolation followed by (e) antigenic subtypes of the HA and NA in parentheses. For example: influenza A/Singapore/1/57 (H2N2) indicates that influenza was first originated from Singapore and was isolated for the first time in the year 1957. The HA and NA antigens are H2 and N2 as shown in the parentheses.

◗ Other properties
Influenza viruses are inactivated by ether, formaldehyde, phe-nol, salts of heavy metals, and common chemical disinfectants. Heating at 50°C for 30 minutes inactivates most strains of the virus. Influenza viruses are resistant to slow drying. They remain viable in dust up to 2 weeks, at 270°C for a longer period, and by freeze-drying indefinitely.
Related Topics