Chapter: Genetics and Molecular Biology: Lambda Phage Genes and Regulatory Circuitry
Induction from the Lysogenic State
Induction from the Lysogenic State
In this section we discuss how lambda escapes from
the lysogenic state and enters a lytic cycle. One time at which this occurs
follows host DNA damage and repair. The signal a phage uses for detection of
this damage is the extensive binding of RecA protein to single-stranded DNA
that accumulates at a replication site when extensive DNA damage is pre-sent.
Apparently when polymerized along the single-stranded DNA, RecA is held in a
shape that interacts with LexA and lambda repressor. The interaction stimulates
the inherent self-cleavage activity of these repressors (Fig. 14.15). Once
cleaved, these repressors are no longer able to repress.
In a nonlysogenic cell the genes that are normally
repressed by LexA protein are the lexA
gene itself, the recA gene, and a set
of about 20 others that are part of the SOS system. Known functions of this
system are to repair damaged DNA and to postpone cell division until repair is
completed. Once repair is completed, the single-stranded DNA no longer
available to activate RecA. Consequently, newly synthesized LexA pro-tein is no
longer cleaved and it therefore represses synthesis of RecA and the other
proteins of the SOS system. In a normal nonlysogenic cell, the SOS system
switches off when it is no longer needed.
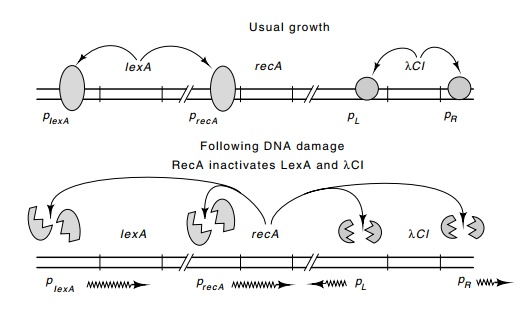
Figure
14.15 LexA protein represses its own
synthesis as well as that of RecAprotein. Cleavage of LexA and lambda repressor
activated by RecA derepresses the lexA,
recA, and lambda operons.
The proteolytic activity of LexA and CI which is
stimulated by RecA protein disconnects the N-terminal domain of the lambda
repressor from the C-terminal domain (Fig. 14.16). A number of physical
experi-ments with protease-digested repressor or with N-terminal fragments of
repressor produced from nonsense mutations have revealed that the N-terminal
half of the protein folds up to form a compact domain that can bind to
operator. The C-terminus also folds to form a compact domain, but this domain
is primarily responsible for the dimerization of repressor. The C-terminal
domains lacking N-terminal domain still dimerize, whereas the N-terminal
fragments do not.
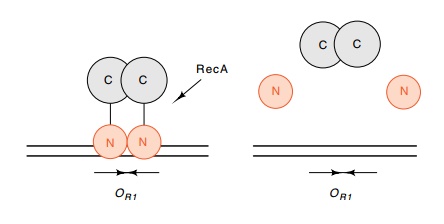
Figure
14.16 Representation of the structure
of lambda repressor and theeffects of cleavage by RecA protease. The
DNA-binding domains are shown contacting one another while on DNA because a
small bit of the dimerization energy derives from such an interaction.
As required for induction,
the affinity of the monomeric N-terminal domain for operator is much lower than
that of the intact dimeric repressor. Furthermore, the N-terminal domains do
not show strong cooperativity in their binding to adjacent operators. In the
next section we will see why the proteolytic cleavage and elimination of
dimeric DNA-binding domains greatly reduce the affinity of repressor for
opera-tor. As a result of this reduced affinity, repressor comes off the DNA and
the phage induces.
The
N-terminal domain of repressor makes protein-protein contacts with RNA
polymerase when it stimulates transcription from the pRM promoter (Fig. 14.17). The stimulation provided by
this positive-acting factor is similar to the stimulation provided by CRP
protein. The repressor accelerates the isomerization rate by RNA polymerase
after it has bound to pRM.
CRP bound near position -41 on promoters also stimulates isomerization.
Figure 14.17 The activation of RNA polymerase atpRMby two repressordimers bound at OR1 and OR2.
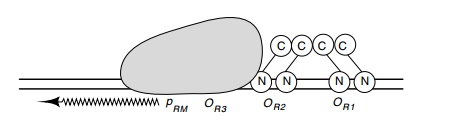
Two types
of evidence indicate that it is the N-terminal domain of repressor that
contacts RNA polymerase. The first is the existence of mutations in lambda
repressor that eliminate the positive stimulation of pRM by repressor when it occupies OR1 and OR2.
These lie in the portion of the protein immediately adjacent to the helices of
the N-ter-minal domain that contact operator. The second is that high levels of
the N-terminal domain are capable of stimulating pRM.
Related Topics