Chapter: Pharmaceutical Biotechnology: Fundamentals and Applications : Oligonucleotides
Improving Oligonucleotide Stability
IMPROVING OLIGONUCLEOTIDE STABILITY
Nuclease resistance of AS-ONs can be improved by modifying the backbone
of the ONs. Since the early 60s of the last century, several chemical
modifications have been introduced to prevent such enzymatic degradation. The
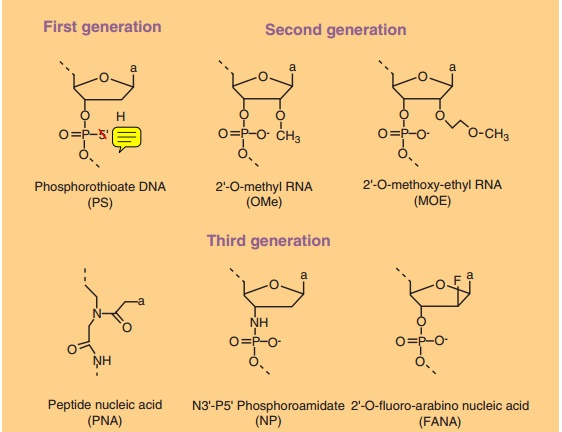
first generation DNA analogs con-sisted of the phosphorothioate (PS) ONs, in
which one of the non-bridging oxygen atoms in the phosphodie-ster bond is replaced
by sulfur (Fig. 1). These PS-ONs are more stable in serum, but also show higher
binding to proteins that than unmodified ONs, which can cause toxicity
problems. However, protein bind-ing also contributes to the increased
circulation half-life seen for PS-ON (40–60 hours), presumably due to reduced
renal excretion. The second generation ONs are those containing alkyl
modifications at the 20 position of the ribose unit
(Fig. 1). Although less toxic than the PS-ONs, they have the disadvantage to be
a poor substrate for RNase H and thus can only inhibit translation by forming a
steric block. However, ONs consisting of 20-O-(2-methoxyethyl)
ONs have greatlyimproved plasma and tissue half-lives (up to 30 days),
presumably due to reduced clearance by nuclease attack. Third generation ONs
consist of a variety of different chemical modifications all with the aim to
improve stability, pharmacokinetics, and interaction with RNA (Fig. 1).
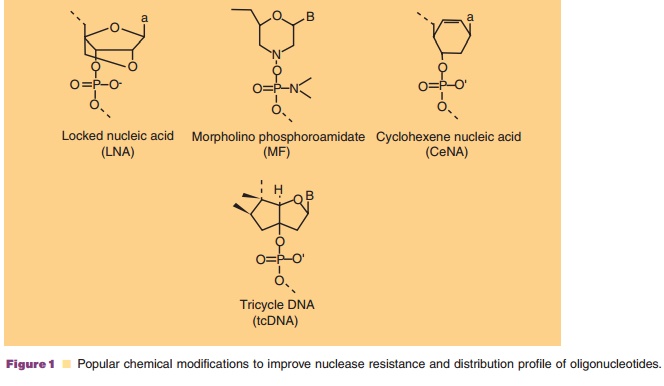
Examples are protein nucleic acids (PNAs) that have a polyamide backbone rather
than a deoxyribose phosphate backbone, locked nucleic acids (LNAs) that have a
methylene bridge between the 20-oxygen
of the ribose and the 4 0-carbon, and morpholino nucleic
acids containing a non-ionic morpholino subunit instead of a ribose, interlinked
by a phosphoroamidate bond (Fig. 1). These third generation ONs all have in
common the superior stability and RNA binding properties, but all lack RNase H
activation. Therefore, chimeras of third generation ONs with DNA have been made
that combine good stability and effective RNase H activation.
Modification of ribozymes or aptamers to obtain better stability and
resistance against nucleolytic degradation is even more challenging, as such
alterations most likely results in loss of respectively enzymatic or binding
activity. The effect of each nucleotide modification on the stability and
activity of the ribozyme or aptamer often have to be established empirically.
The serum half-life of a DNA ribozyme could be 10-fold increased by protecting
the 30-end with inverted nucleotides (Sun et al., 1999).
Stability of siRNA in cell culture and serum is generally not a limiting
factor due to thermal instability of RNase A. However, for protection against
intracel-lular nucleases further enhancement of stability might be required.
This can be achieved by inserting modified nucleotides on either ends of the
siRNA. However, one needs to be aware that the amount and chemical nature of
modified nucleotides may influence the gene silencing efficacy of siRNA
(Amarzguioui et al., 2003). siRNA is generally rapidly cleared.
Related Topics