Chapter: Pharmaceutical Biotechnology: Fundamentals and Applications : Oligonucleotides
Direct Binding to Non-Nucleic Acids
DIRECT BINDING TO NON-NUCLEIC ACIDS
The ability of nucleic acids to fold into complex three-dimensional structures through regions of (partial) complementarity allows them to bind to virtually any molecule with nano- to picomolar affinity (Proske et al., 2005). This high affinity is supported by data on their extreme specificity. A nucleic acid sequence specifically binding theophylline has a million times higher affinity for theophylline than caffeine, mole-cules which differ by only one methyl group.
Aptamers/Riboswitches
Aptamers and riboswitches are single-stranded ONs of either DNA or RNA, generally about 60 nucleotides long, which fold into well-defined three-dimensional structures. They bind to their target molecule by complementary shape interactions accompanied by charge and hydrophobic interactions and hydrogen bridges. The target can be small molecules or macromolecules. Aptamers are isolated artificially, whereas riboswitches occur naturally (Tucker and Breaker, 2005). Several viruses have been shown to encode small, structured RNAs that bind to viral or cellular proteins with high affinity and specificity. It was demonstrated that these RNAs could modulate the activity of proteins essential for viral replication or inhibit the activity of proteins involved in cellular antiviral responses. Also the genomes of prokaryotes have been shown to contain nutrient responsive riboswitches to regulate gene expression.
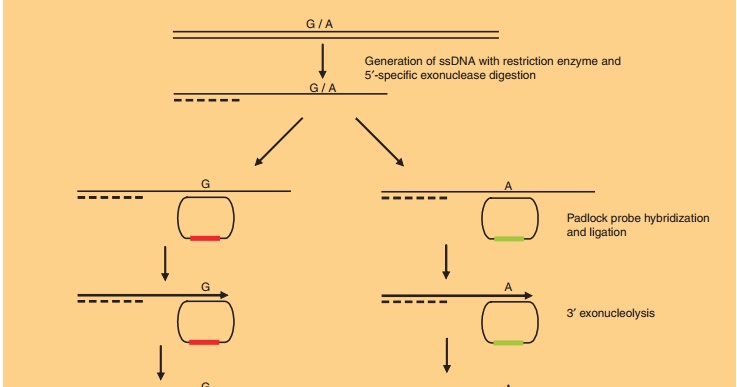
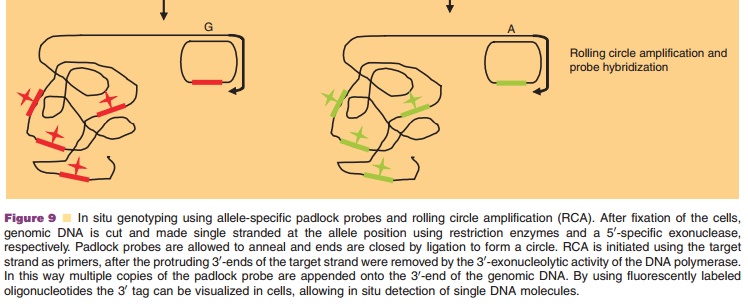
Synthetically, such compounds can be identified by subjecting large libraries of nucleic acid molecules to a panning procedure (Fig. 9). This selection process has been named systematic evolution of ligands by exponential enrichment (SELEX). The resulting li-gands are called aptamers. The SELEX process starts by generating a large library of randomized RNA sequences. This library contains up to 1015 different nucleic acid molecules that fold into different struc-tures depending on their sequence. The library is incubated with the structure of interest, and those RNAs present in the library that bind the protein are separated from those that do not. The obtained RNAs are then amplified by reverse transcriptase-PCR and in vitro transcribed to generate a pool of RNAs that have been enriched for those that bind the target of interest. This selection and amplification process is repeated (usually 8–12 rounds) under increasingly stringent binding conditions to promote Darwinian selection until the RNA ligands with the highest affinity for the target protein are isolated. This molecular evolution process can also be performed with DNA, circumventing the need for reverse transcription before PCR and in vitro transcription. Automation has reduced aptamer in vitro selectiontimes from months to days, making aptamers suitable for application in high throughput target validation. Aptamers can bind to proteins for therapeutic use, like antibodies do. However, antibody selection requires a biological system. The selection of aptamers is a chemical process, therefore it can target almost any protein. In contrast to antibodies, aptamers are prone to bind to functional domains of the target protein (for reasons unknown), such as substrate binding pockets or allosteric sites, thereby modulating the biological function of the molecule. A common problem upon the therapeutic development of apta-mers is that they can be so specific for the human version of a target protein that they have poor cross-reactivity with orthologs of the target from other animal species, making their pre-clinical evaluation difficult. Toggle SELEX is a possible solution in which selection occurs alternating between pre-clinical and clinical target (White et al., 2001).
One aptamer targeting VEGF, pegaptanib (Macugen ), is marketed for wet age-related macular degeneration (Ng et al., 2006). The PEGylated aptamer is injected in the vitreous at a dose of 1.65 mg (0.3 mg of which is aptamer)/eye in 90 ml every 6 weeks. Adverse effects included endophtalmitis and bleeding events in the eye, likely related to the injection procedure. The compound also appeared to exhibit effects in diabetic retinopathy. To increase stability several nucleotides are 20-O-methyl and 20-O-fluoro modified and the aptamer is conjugated to polyethylene glycol, which stabilizes siRNA in solution and facilitates clinical delivery.
Related Topics