Chapter: Pharmaceutical Biotechnology: Fundamentals and Applications : Oligonucleotides
Antisense/Ribozymes/EGS - Oligonucleotides Interfering With Gene Expression
Antisense/Ribozymes/EGS
The function of ONs to act as antisense molecules was discovered by
Zamecnik and Stephenson in 1978, making it the oldest oligonucleotide-based
therapeutic approach (Stephenson and Zamecnik, 1978; Zamecnik and Stephenson,
1978). Many of the difficulties associated with the use of ONs for medical
applications have consequently been encountered for antisense first, explaining
why clinical progress has been difficult. Improvements in synthetic chemistry,
knowledge of genome, transcriptome, and proteome, and delivery issues have
revived interest in the
technology (Chan et al., 2006). “Classical” antisense ONs are
single-stranded DNA or RNA molecules that generally consist of 13 to 25
nucleotides. They are complementary to a sense mRNA sequence and can hybridize
to it through Watson–Crick base-pairing. Three classes of
translation-inhibiting ONs can be distinguished based on their mechanism of
action:
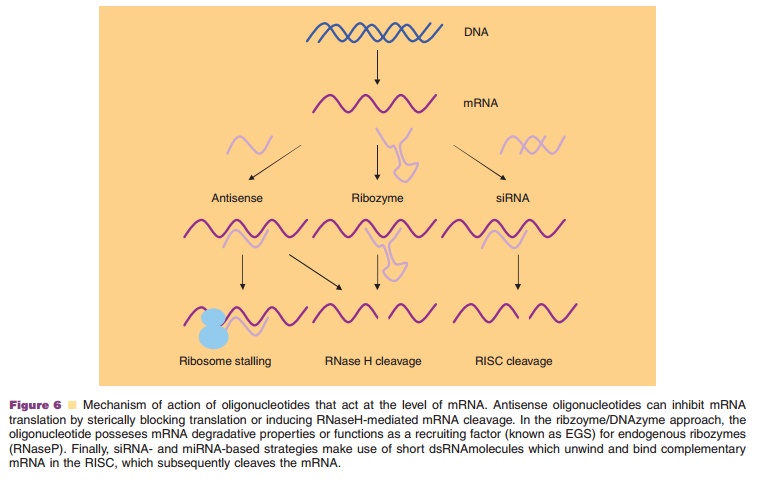
mRNA-blocking ONs, which physically prevent or inhibit the progression
of splicing or translation through binding of complementary mRNA se-quences
(Fig. 6).
mRNA cleaving ONs, which induce degradation of mRNA by binding
complementary mRNA sequences and recruiting the cytoplasmic nucleaseRNase H
(Aboul-Fadl, 2005).
mRNA cleaving ONs, which induce degradation of mRNA by recruiting
nuclear RNase P via EGS, or by nuclease activity of the nucleic acid itself
(ribozymes/DNAzymes) (Fig. 7) (Lilley, 2005).
The majority of the clinically studied antisense ONs act through RNase
H. RNase H-mediated knockdown generally reaches >80%
downregulation of protein and mRNA expression. In contrast to blocking ONs,
RNase H-recruiting antisense ONs can inhibit protein expression without a
priori restrictions to the region of the mRNA that is targeted. Most blocking
ONs, however, require targeting regions within the 50-untranslated region or AUG initiation codon region as the ribosome is
apparently able to remove bound antisense molecules in the coding region.
One antisense drug has so far been introduced to the market: Fomivirsen
(Vitravene ), for the treatment of cytomegalovirus-induced retinitis in AIDS
patients (Jabs and Griffiths, 2002). Vitravene is injected into the vitreous at
a dose of 165 mg or 330 mg/eye in 25 ml, once weekly for 3 weeks,
followed by 2 weekly administrations. Reported side effects are related to
irritation and inflammation of the eye likely caused by the injection
procedure. The ON has a thioate back-bone which limits nuclease degradation.
Local injec-tion at the pathological site improves target cell accumulation.
This successful introduction is overshadowed by failures in phase III
clinical trials of a number of products in recent years. Alicaforsen did not
induce clinical remissions compared to placebo in patients with Crohn’s
disease, and AffinitakTM (targeting protein kinase C-alpha) and oblimersen (Genasense ; Bcl-2
antisense oligonucleotide) failed to substantially prolong survival in cancer
patients. Nevertheless, the experiences from these trials have provided new
insights into the use of alicaforsen and Genasense , and consequently, these
compounds are still in clinical trials for one or more specific indications.
Table 1
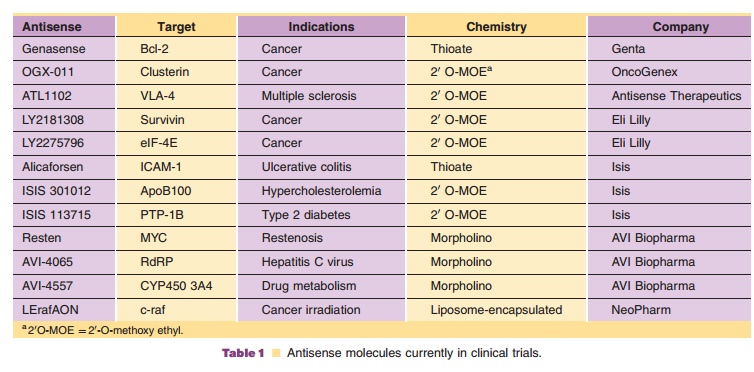
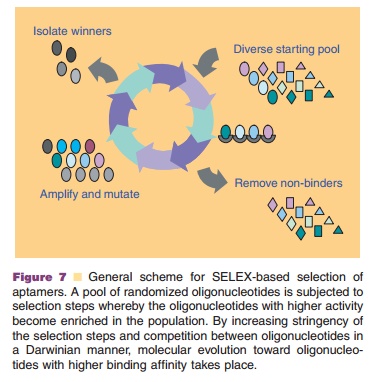
Although sequence specificity is one of the most attractive features for
antisense application, there are reports that show that knockdown of related
genes with only limited sequence homology can occur. In addition, the effects
of oblimersen appear only partly due to Bcl-2 downregulation but can also be
attributed to immune stimulation and mitochondrial apoptosis activation
independent of Bcl-2 (Gekeler et al., 2006; Lai et al., 2006).
Ribozymes and DNAzymes are molecules that combine a mRNA binding sequence with a catalytic domain and are capable of cleaving mRNA molecules. They are potentially capable of multiple turnovers. Several different types of ribozymes are found in nature: the hammerhead, hairpin, hepatitis delta virus (HDV), Varkud satellite RNA, group I and II introns and the RNA subunit of RNase P. Smaller ribozymes, like hammerhead and hairpin, consist of 40 to 150 nucleotides. But other ribozymes can be hundreds of nucleotides long and fold into protein-like structures ontaining single- and double-stranded regions, base-triplets, loops, bulges, and junctions (like RNaseP).
Ribozymes have been applied clinically. In a study in HIV patients,
hematopoietic progenitor cells, were transduced ex vivo using a retroviral
vector carrying an anti-HIV-1 ribozyme. Sustained output of ribozyme expressing
mature myeloid and T-lymphoid cells was detected, showing that the concept may
work (Macpherson et al., 2005).
DNAzymes structurally and functionally resem-ble ribozymes, but are made
of DNA. They are artificial molecules and have, so far, not been found in
nature. The working mechanism of DNAzymes is similar tothat of ribozymes, but
they offer some advantages. Because they are made of DNA, they are easier and
less expensive to synthesize, they are much more resistant to degradation, and
possess improved catalytic effi-ciency. DNAzyme recently produced therapeutic
ef-fects in animal models of ischemia, inflammation and cancer (Fiammengo and
Jaschke, 2005).
RNaseP is an endogenous nuclear ribozyme that is substantially larger
(several hundred bases) than hammerhead and hairpin ribozymes, which makes it
far more difficult to apply exogenously (Trang et al., 2004). However, by
making use of ONs that function as so-called small EGS, they can together with
the mRNA form a structure that resembles the endogen-ous target of RNaseP and
thereby recruits the enzyme to digest the mRNA-oligonucleotide sequence combi-nation
(Tafech et al., 2006). This concept has not yet been validated in vivo.
Related Topics