Chapter: Microbiology and Immunology: Bacteriology: Salmonella
Epidemiology - Salmonella
Epidemiology
◗ Geographical distribution
Enteric fever is endemic in many developing countries of the world and is primarily found in those countries of the develop-ing world where sanitary conditions are poor.
Typhoid fevers are endemic in the Indian subcontinent, Southeast and Far East Asia, the Middle East, Africa, Central America, and South America. Approximately, 12–13 million cases of typhoid fever occur globally each year with 600,000 deaths. Enteric fever is endemic in all parts of India. Typhoid fever has been virtually eliminated in the developed countries due to improvements in water supply and sanitation during last many decades.
Paratyphoid fever still continues to be health problem in both developing and developed countries. S. Paratyphi A is prevalent in India and other Asian countries, Eastern Europe, and South America; S. Paratyphi B in North America, Britain, and Western Europe; and S.Paratyphi C in Eastern Europe and Guyana.
◗ Habitat
S. Typhi and S. Paratyphi (A, B, and C) are strict human patho-gens. They are not found in any other animal hosts. They colonize the small intestine, especially ileac mucosa in infected human hosts. Asymptomatic long-term colonization occurs commonly in infected hosts.
Other salmonellae are parasitic in various domestic animals, rodents, reptiles, and birds. S. Typhimurium have a wide host range affecting animals, birds, and humans, while Salmonella Abortus-equi is found only in horses, Salmonella Abortus-oris in sheep, andS.Gallinarum in poultry.
◗ Reservoir, source, and transmission of infection
The infected patient and, more frequently, carriers are impor-tant reservoirs of infection for enteric fever.
About 2–4% of patients become chronic carriers. The bacilli persist in the gallbladder and are excreted in feces (fecal carrier) or persist in the kidney and are secreted in the urine (urinarycarrier). Urinary carriers are less frequent than the fecal carriersand usually present with some urinary lesion, such as calculus or schistosomiasis.
The carrier state is more common in women than in men and in the older people (over 40 years) than in young people.
Food handlers or cooks who become carriers are potentially dangerous to the community. Mary Mallon (Typhoid Mary), a New York cook was a classical case of such carrier who, over a period of 15 years, caused at least seven outbreaks affecting more than 200 persons.
Carrier stage also occurs with S. Paratyphi infections. S. Paratyphi A infection occurs only in humans, while S. Paratyphi B infection occurs in animals, such as dogs or cows.
Food, vegetables, and water contaminated with human feces infected by S. Typhi are the common sources of infection.
· S. Typhi infections occur when food or water contaminatedby infected food handlers or due to poor personal hygiene is ingested.
· The infectious dose for S. Typhi infections is low, so person-to-person spread is common.
· The infectious dose is still lower for people at high risk for disease because of age, immunosuppression, or underlying disease (leukemia, lymphoma, sickle cell disease), or reduced gastric acidity.
There is no animal reservoir for typhoidal salmonellae. Animal-to-human zoonotic transmission is common in only nonty-phoidal salmonellae. Poultry, livestock, reptiles, and pets are the principal reservoirs for nontyphoidal Salmonella organisms. Ingestion of improperly cooked fruits, vegetables, foods of ani-mal origin, including poultry, red meats, unpasteurized milk, and eggs that have been contaminated by infected animals or an infected human is the mode of transmission. Contact with infected reptiles, such as iguanas, pet turtles, and tor-toises, and ingestion of contaminated water are other modes of transmission.
◗ Classification and typing of Salmonella
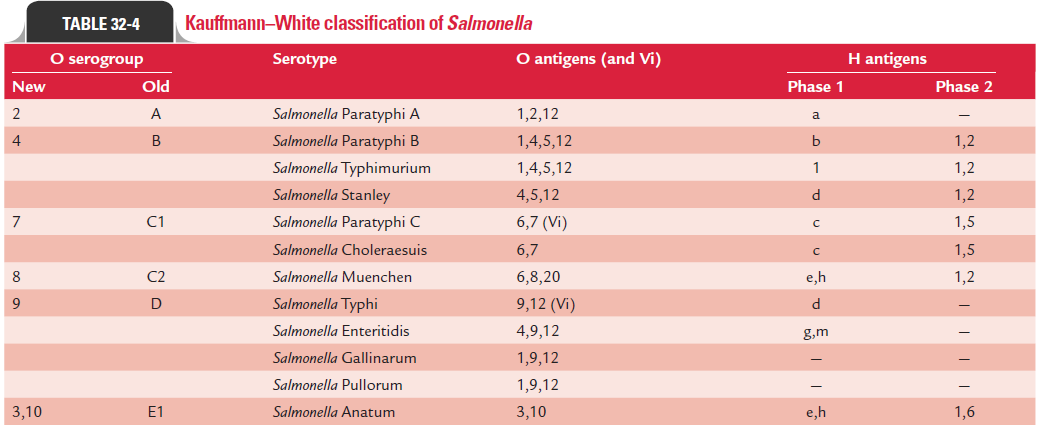
1. Kauffmann–White scheme: Kauffmann–White schemeis a method of classification of Salmonella strains based on structural formulae of the O and H antigens of the strains, which are identified by agglutination with specific anti-sera. Salmonellae are classified into different O serogroups, based on the presence of distinctive O antigen factors. Each serogroup contains a number of serotypes possessing a common O factor not found in other serogroups. These O factors are now designated 1, 2, 3, 4, etc.; these were originally designated by capital letters A–Z and afterward by numbers 51–67. Some serogroups were subdivided into subserogroups (e.g., C1–C4; E1–E4). It seems logical to name each serogroup by its characteristic O antigen factor numbers, rather than by letters. Hence, it is now being pro-posed to designate group A as 2, B as 4, C1 as 7, C2 as 8, D as 9, and so on.
Within each O serogroup, the different serotypes are identified by the presence of phase 1 and phase 2 of flagellar antigens. The antigenic structure of a Salmonella serotype is designated by an antigenic formula, which has three components describing the O antigens, the phase 1 H antigen, and phase 2 H antigen in that order. Three parts are separated by colons and the component antigens in each part by commas.
The Kauffmann–White scheme designates each serotype as a species (Table 32-4). Currently, more than 2400 sero-types have been described by Kauffmann–White scheme.
2. Bacteriophage typing: Bacteriophage typing is the intra-species classification of S. Typhi, first developed by Craigie and Yen (1937). They observed that bacteriophage acting on the Vi antigen of S. Typhi (Vi phage II) is highly adapt-able. The parent phage is known as phage A. It could be made specific for a particular strain of S. Typhi by serial propagation in the strain. Such adaptation was obtained by phenotypic or genotypic variation.
The phage typing is carried out by determining the sensitivity of the cultures to a series of variants of Vi phase. However, most other types are sensitive to only one or a few related adaptions. As phage typing of S. Typhi depends on the presence of Vi antigens, a proportion of strains (Vi negative) will be untypable. At present, 97 Vi II phage types of S. Typhi are recognized. S. Typhi phage types E1, O, and A are most common in India.
To make phage typing more useful and more discrimi-native, additional markers have been used for the subdivi-sion of strains belonging to a phage type. These include (a) Nicolle’s complementary phage typing, (b) Kristensen’s biotyping, (c) production of tetrathionate reductase, (d) production of bacteriocin, and (e) antibiogram. The phage typing is useful in (i) tracing the source of epidemics and (ii) providing information on the trends and patterns of typhoid epidemiology at the local, national, and interna-tional levels. Phage typing schemes have also been applied to S. Paratyphi A and B, S. Typhimurium, S.nteritidis, S. Dublin, and other serotypes. Phage typing in India is car-ried out at the National Phage Typing Centres located at the Lady Hardinge Medical College, New Delhi.
3. Biotyping: Biotyping is a useful method to classify differ-ent Salmonella strains of a particular serotype into a num-ber of biotypes by their biochemical characteristics.
On the basis of 15 different biochemical characteristics, Duguid et al. in 1975 differentiated S. Typhimurium into 144 different biotypes. The same biochemical characteris-tics have also been used to biotype S. Paratyphi B, Salmonella Montevideo, and S.Agona. S. Typhi has been differenti-ated into different biotypes on the basis of fermentation of arabinose, dulcitol, and xylose.
Biotyping is useful to supplement phage typing because it subdivides further a large number of untypable strains or members of common phage types. Conversely, strains of the same biotype may be reorganized into different phage types. Therefore, a combination of phage typing and biotyping provides a better discrimination of Salmonella strains than either method used alone.
4. Molecular methods: Currently, genotyping methods likeplasmid fingerprinting, multilocus enzyme electrophore-sis, IS-200 profiling, and random amplified polymorphic DNA analysis have been employed as more discriminating methods for epidemiological characterization of Salmonella in advanced centers.
The National Salmonella Reference Centre in India is located at the Central Research Institute, Kasauli (Himachal Pradesh). The reference center for salmonel-lae of animal origin is at the Indian Veterinary Research Institute, Izatnagar (UP). Theses reference centers provide services for identification of unusual serotypes and recon-firmation of other serotypes of salmonellae.
Related Topics