Chapter: Genetics and Molecular Biology: Induction, Repression, and the araBAD Operon
DNA Looping and Repression of araBAD
DNA Looping and Repression of araBAD
How can
repression of pBAD be
generated from the araO2
site that is located more than 200 base pairs upstream? There are three
possibilities (Fig. 12.11). A signal could be transmitted through the DNA, for
exam-ple, by changing the angle of the base tilt; something could polymerize
along the DNA starting at araO2
and finally cover araI; or the DNA
could bend back and bring the araO2
region near pBAD. This
latter possibility was shown to be the case by a series of experiments in which
the spacing between araO2
and the promoter region was varied. If five base pairs, which is half a helical
turn, are added between these two sites, the ability to repress is greatly
diminished, but if 11 base pairs are added, the ability to repress is restored.
An addition of 15 base pairs eliminates repression, and a longer addition of 31
base pairs restores repression.
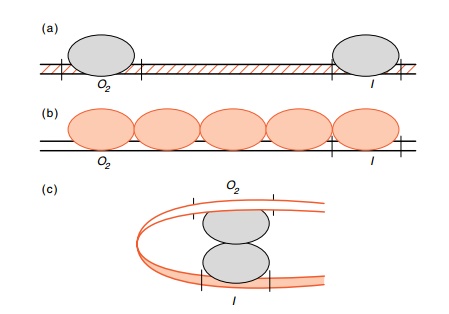
Figure
12.11 Three possible mechanisms for
repression from a site severalhundred nucleotides from a promoter (a)
properties of the DNA can be altered, (b)
protein can polymerize along the DNA, or (c) loops can form.
These results show that the absolute distance between the two sites is not greatly important but that the angular orientation between the two is.
This implies that the DNA loops back to form contacts between AraC protein
bound at araO2 and
something in the promoter region. If the protein at araO2 is on the opposite side of the DNA (Fig. 12.12),
the additional energy required to twist the DNA half a turn so the sites again
face one another is too great and the loop required for repression does not
form. The isolation of repression deficient point mutations located the sites
necessary for repression. As expected, some were in araO2. Additional repression negative mutations were in araI, thereby identi-fying the other end
of the loop.
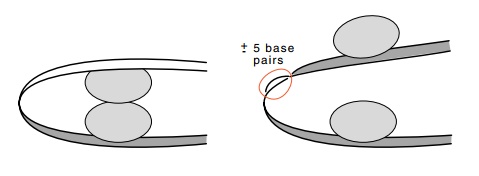
Figure
12.12 How a five-base-pair insertion
can introduce half a helical turnand prevent formation of a loop.
Related Topics