Chapter: Plant Biochemistry: Biotechnology alters plants to meet requirements of agriculture, nutrition and industry
Agrobacteria can transform plant cells
Agrobacteria can transform plant cells
Gram-negative soil bacteria of the species Agrobacterium tumefaciens (new nomenclature: Rhizobium radiobacter) induce a tumor growth at wounding sites in various plants, often on the stem, which can lead to the formation of crown galls. The tumor tissue from these galls continues to grow as a callus in cell culture. Mature differentiated plant cells, which normally no longer divide, can be stimulated to unrestricted growth by the addition of the phytohormones auxin and cytokinin. In this way a tumor in the form of a callus can be obtained from a differentiated plant cell. The gall produced by theAgrobacterium is formed in basically the same way. The bacteria force the wounded plants to produce high con-centrations of auxin and cytokinin, resulting in a proliferation of the plant tissue and leading to tumor growth. Since the capacity for increased cytoki-nin and auxin synthesis is inherited after cell division by all the succeeding cells, a callus culture of crown gall tissue can be multiplied without adding phytohormones.
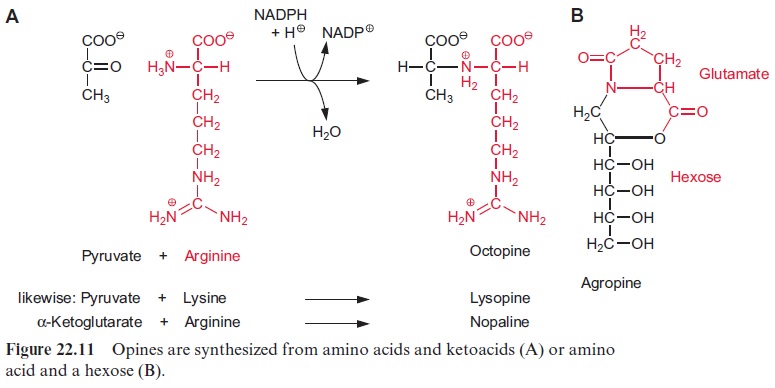
The crown gall tumor cells have acquired yet another ability: they can produce a variety of products named opines by condensating amino acids and -ketoacids or amino acids and sugars. These opines are synthesized in such high amounts that they are excreted from the crown galls. Figure 22.11 shows octopine, lysopine, nopaline, and agropine as examples ofopines. Each Agrobacterium strain induces the synthesis of only a single opine. The synthesis of the first three opines mentioned proceeds via con-densation to form a Schiff base and a subsequent reduction by NADPH. The opines are so stable that they cannot be metabolized by most soil bac-teria. Agrobacterium tumefaciens strains have specialized in utilizing these opines. Normally, a particular opine can be catabolized only by that bacte-rial strain which has induced its synthesis in the plant. In such a way the wounded plants are forced to produce a special nutrient that can be con-sumed only by the corresponding Agrobacterium.
The conversion of differentiated plant cells to opine-producing tumor cells occurs without the Agrobacteria entering those cells. This reprogram-ming of plant cells is caused by the transfer of functional genes from the bacteria to the genome of the wounded plant cells. The Agrobacteria have acquired the ability to transform plants in order to use them as a produc-tion site for their nutrient. Jeff Schell named this novel parasitism genetic colonization.
The Ti plasmid contains the genetic information for tumor formation
The phytopathogenic function of A. tumefaciens is encoded in a tumor-inducing Ti plasmid with a size of about 200 kbp (Fig. 22.12). Strains of A. tumefaciens containing no Ti plasmids are unable to induce the forma-tion of crown galls.
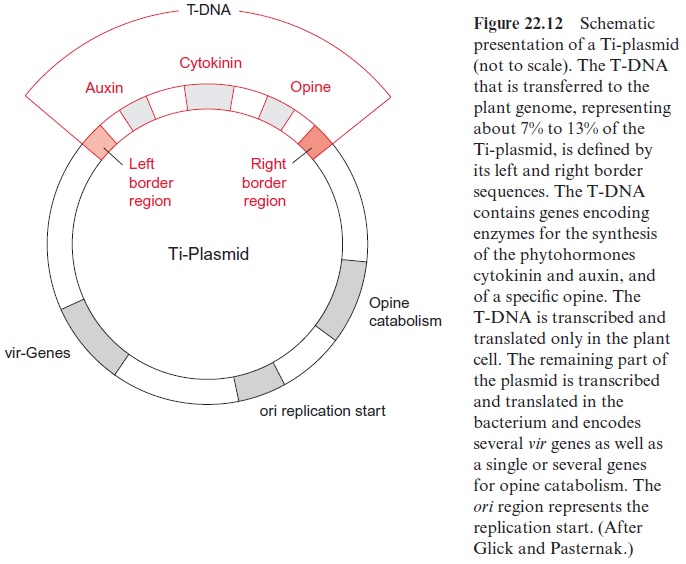
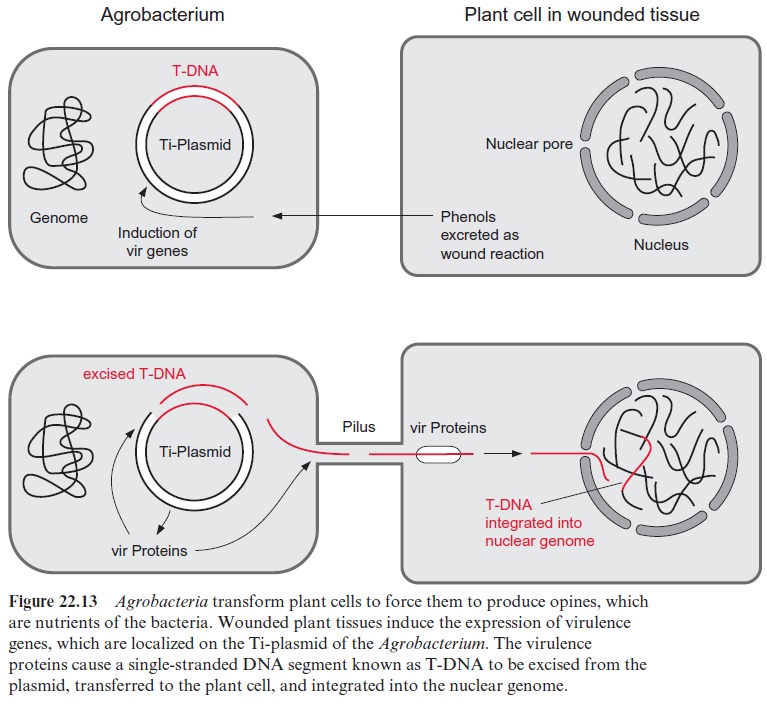
Plants are infected by bacteria at wounds frequently occurring at the stem base. After being wounded, plants excrete phenolic compounds as a defense against pathogens . These phenols are used by the agrobacteria for the chemotactic localization of the wounded plant cells and the subsequent initiation of infection. Phenols stimulate the expression of about 11 virulence genes (vir genes) located on the Ti plasmid. These vir genes encode virulence proteins which enable the transfer of the bacte-rial tumor-inducing genes to the plant genome. From a 12 to 25 kbp long section of the Ti plasmid, named T-DNA (T, transfer), a single strand is excised by a vir nuclease. The cleavage sites are defined by border sequences present at both ends of the T-DNA. The transfer of the single strand T-DNA from the bacterium to the plant cell nucleus (Fig. 22.13) proceeds in analogy to the conjugation, which is a bacterial sexual reproduction process. To connect the bacterium to the plant cell, vir proteins form a pilus, a threadlike structure through which the T-DNA is conducted. After being transferred into the plant cell, the T-DNA strand migrates further into the nucleus. Vir-encoded proteins protect the T-DNA from being attacked en route by DNA-degrading plant enzymes and also facilitate the transport through the nuclear pores to the nucleus. The right border region of the T-DNA has an important function in its integration into the plant nuclear genome. The T-DNA is integrated randomly into chromosomes and, when inserted in a gene, can eliminate the function of this gene. This can be used to identify a gene by gene tagging in an analogous way as the gene tagging by the transposons.
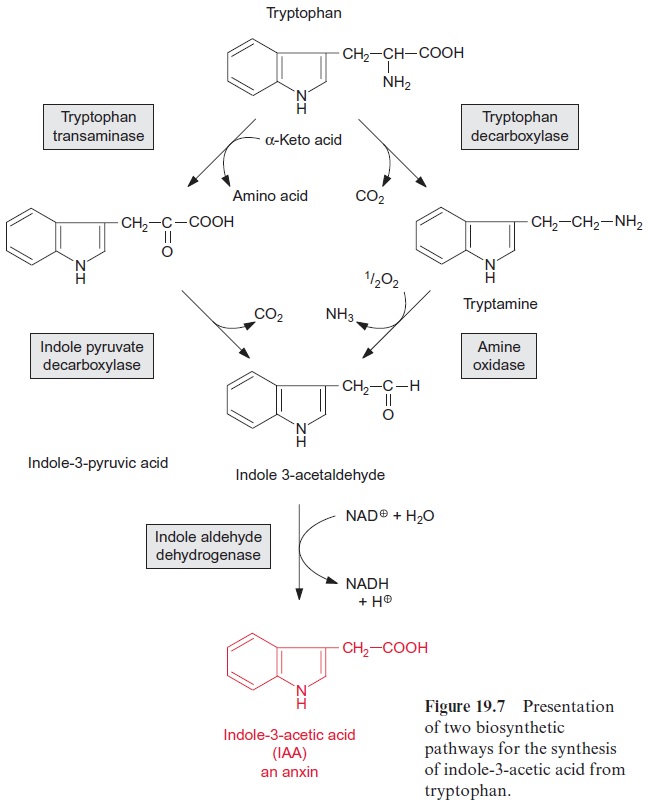
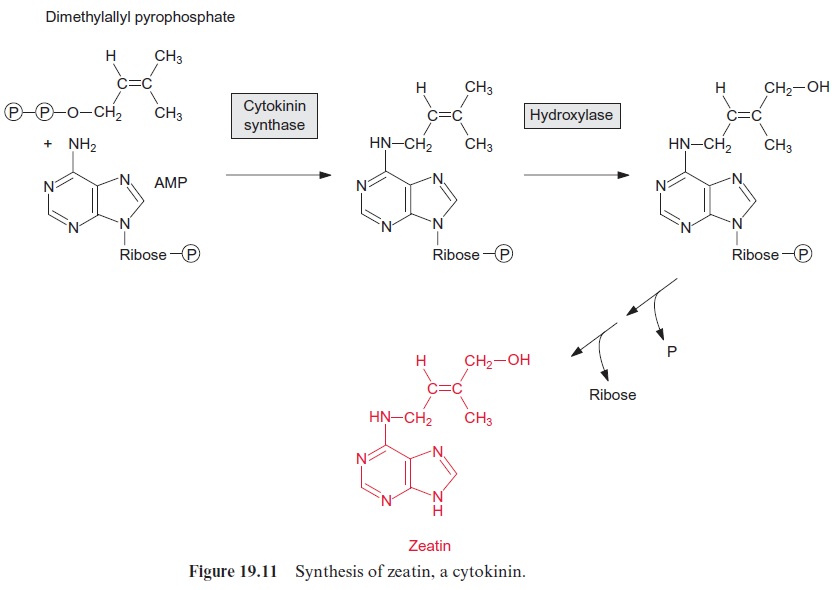
The T-DNA integrated in the nuclear genome has the properties of a eukaryotic gene and is inherited in a Mendelian fashion. It is replicated by the plant cell as if it were its own DNA and, since it contains eukaryo-tic promoters, is also transcribed. The resultant mRNA corresponds to a eukaryotic mRNA and is translated as such. The T-DNA encodes cytoki-nin synthase, a key enzyme for the synthesis of the cytokinin zeatin (Fig. 19.11), as well as two enzymes for the synthesis of the auxin indoleacetic acid (IAA). This bacterial IAA synthesis proceeds in a different manner than plant IAA biosynthesis (Fig. 19.7), but details of this pathway will not be considered here. Moreover, the T-DNA encodes one or two enzymes, varying from strain to strain, for the synthesis of a special opine.
The enzymes required for catabolism of the corresponding opine are encoded in that part of the Ti plasmid that remains in the bacterium. Thus, in parallel to the transformation of the plant cell, the enzymes for opine degradation are synthesized by the bacteria.
Related Topics