DNA and RNA | Formation, Structure, Features, Types - Nucleic Acids - Biomolecules | 11th Botany : Chapter 8 : Biomolecules
Chapter: 11th Botany : Chapter 8 : Biomolecules
Nucleic Acids - Biomolecules
Nucleic Acids
As we know DNA and RNA are the two kinds of nucleic
acids. These were originally isolated from cell nucleus. They are present in
all known cells and viruses with special coded genetic
programme with detailed and specific instructions for each organism heredity.
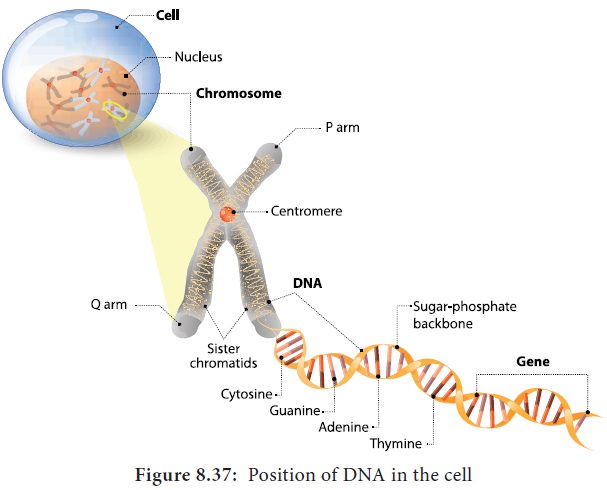
DNA and RNA are polymers of monomers called nucleotides, each of which is composed
of a nitrogen base, a pentose sugar and a phosphate. A purine or a pyrimidine
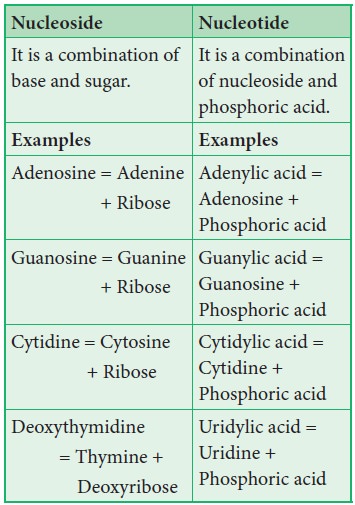
and a ribose or deoxyribose sugar is called nucleoside. A nitrogenous base is linked to pentose sugar through
n-glycosidic linkage and forms a nucleoside. When a phosphate group is attached
to a nucleoside it is called a nucleotide.
The nitrogen base is a heterocyclic compound that can be either a purine (two rings) or a pyrimidine (one ring). There are 2 types of purines – adenine (A) and
guanine (G) and 3 types of pyrimidines – cytosine (C), thymine (T) and uracil
(U) (Figure 8.38).
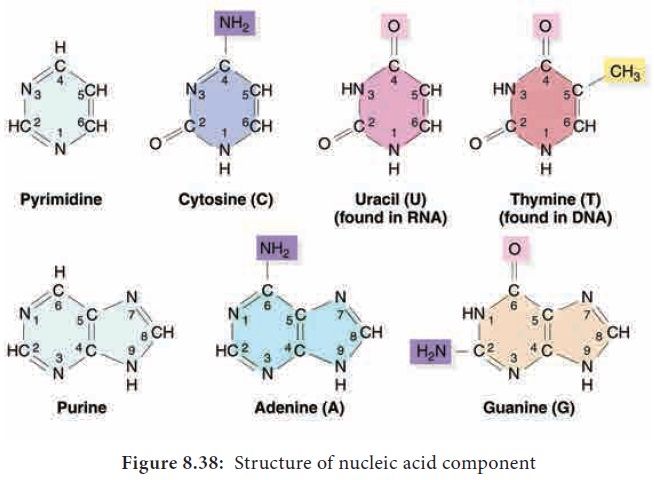
A characteristic feature that
differentiates DNA from RNA is that DNA contains nitrogen bases such as
Adenine, guanine, thymine (5-methyl uracil) and cytosine and the RNA contains nitrogen
bases such as adenine, guanine, cytosine and uracil instead of thymine. The
nitrogen base is covalently bonded to the sugar ribose in RNA and to
deoxyribose (ribose with one oxygen removed from C2) in DNA.
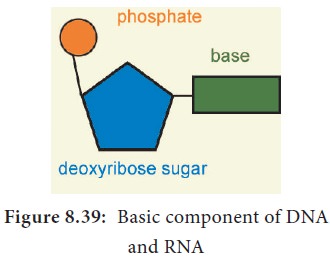
Phosphate group is a derivative of (PO43-) phosphoric acid, and
forms phosphodiester linkages with sugar molecule (Figure 8.39).
1. Formation of Dinucleotide and Polynucleotide
Two nucleotides join to form dinucleotide that are linked through 3′-5′ phosphodiester
linkage by condensation between phosphate groups of one with sugar of other.
This is repeated many times to make polynucleotide.
2. Structure of DNA
Watson and Crick shared the Nobel Prize in 1962 for
their discovery, along with Maurice
Wilkins, who had produced the crystallographic
data supporting the model. Rosalind
Franklin (1920–1958) had earlier produced
the first clear crystallographic evidence for a helical structure. James Watson and Francis Crick (Figure
8.40) of Cavendish laboratory in
Cambridge built a scale model of double helical structure of DNA which is the
most prevalent form of DNA, the B-DNA.
This is the secondary structure of DNA.

As proposed by James
Watson and Francis Crick, DNA
consists of right handed double
helix with 2 helical polynucleotide chains that are coiled around a common axis
to form right handed B form of DNA. The coils are held together by hydrogen
bonds which occur between complementary pairs of nitrogenous bases. The sugar
is called 2′-deoxyribose
because
there is no hydroxyl at position 2′. Adenine and thiamine base pairs
has two hydrogen bonds while guanine and cytosine base pairs have three
hydrogen bonds.
Chargaff
’s Rule:
•
A = T; G ≡ C
•
A + G = T + C
•
A : T = G : C = 1
As published by Erwin Chargaff in 1949, a purine pairs with pyrimidine and vice
versa. Adenine (A) always pairs with
Thymine (T) by double bond and Guanine (G) always pairs with Cytosine (C) by
triple bond.

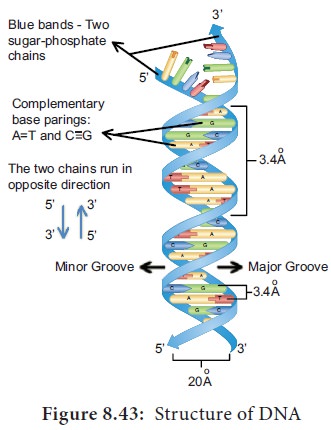
3. Features of DNA
• If one strand runs in the 5′-3′ direction, the other runs in 3′-5′ direction and thus are antiparallel (they run in opposite direction). The 5′ end has the phosphate group and 3’end has the OH group.
•
The angle at which the two sugars protrude from the
base pairs is about 120°, for the
narrow angle and 240° for the
wide angle. The narrow angle between the sugars generates a minor groove and the large angle on the other edge generates major
groove.
•
Each base is 0.34 nm apart and a complete turn of
the helix comprises 3.4 nm or 10 base pairs per turn in the predominant B form
of DNA.
•
DNA helical structure has a diameter of 20 A° and a pitch of about 34 A°. X-ray crystal study of DNA
takes a stack of about 10 bp to go completely around the helix (360°).
•
Thermodynamic stability of the helix and
specificity of base pairing includes (i)the hydrogen bonds between the
complementary bases of the double helix (ii) stacking interaction between bases
tend to stack about each other perpendicular to the direction of helical axis.
Electron cloud interactions (∏ – ∏) between the bases in the
helical stacks contribute to the stability of the double helix.
•
The phosphodiester linkages gives an inherent
polarity to the DNA helix. They form strong covalent bonds, gives the strength
and stability to the polynucleotide chain (Figure 8.43).
•
Plectonemic coiling - the two strands of the DNA
are wrapped around each other in a helix, making it impossible to simply move
them apart without breaking the entire structure. Whereas in paranemic coiling
the two strands simply lie alongside one another, making them easier to pull
apart.
•
![]()
![]()
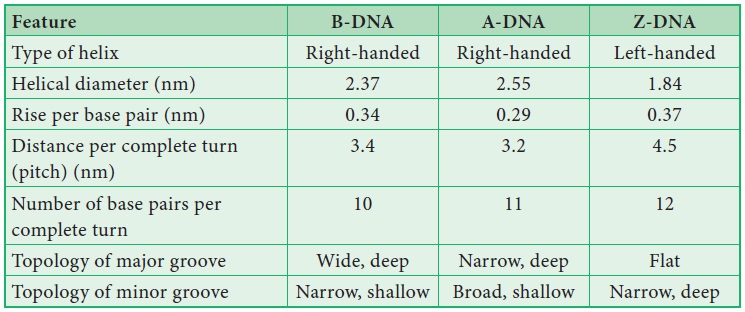
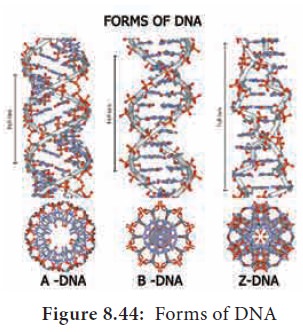
![]() Based on the helix and the
distance between each turns, the DNA is of three forms – A DNA, B DNA and Z DNA (Figure
8.43) .
Based on the helix and the
distance between each turns, the DNA is of three forms – A DNA, B DNA and Z DNA (Figure
8.43) .
4. Ribonucleic Acid (RNA)
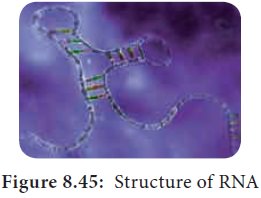
Ribonucleic
acid (RNA) is a polymeric molecule
essential in various biological roles in coding, decoding, regulation and
expression of genes. RNA is single stranded and is unstable when compared to
DNA (Figure 8.45).
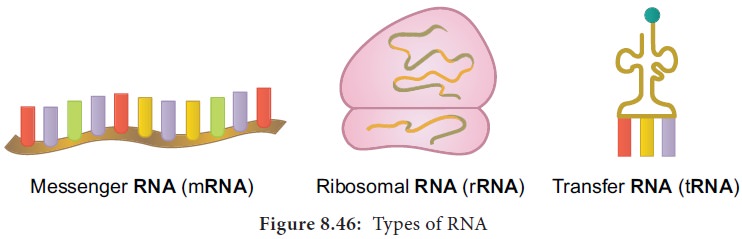
5. Types of RNA
·
mRNA
(messenger RNA): Single stranded, carries a copy of instructions for
assembling amino acids into proteins. It is very unstable and comprises 5% of total RNA polymer.
Prokaryotic mRNA (Polycistronic) carry coding sequences for many polypeptides.
Eukaryotic mRNA (Monocistronic) contains information for only one polypeptide.
·
tRNA
(transfer RNA): Translates the
code from mRNA and transfers amino acids to the ribosome to build proteins.
It is highly folded into an elaborate 3D structure and comprises about 15% of total RNA. It is also called
as soluble RNA.
·
rRNA
(ribosomal RNA): Single stranded,
metabolically stable, make up the two subunits of ribosomes. It constitutes 80% of the total RNA. It is a
polymer with varied length from 120–3000 nucleotides and gives ribosomes their
shape. Genes for rRNA are highly conserved and employed for phylogenetic
studies (Figure 8.46).
Related Topics