Chapter: Biotechnology Applying the Genetic Revolution: Environmental Biotechnology
Natural Attenuation of Pollutants
NATURAL
ATTENUATION OF POLLUTANTS
Metagenomics can also be
applied to biogeochemical research. Perhaps identifying how bacteria affect the
environment will help us figure out ways to maintain our species. Understanding
how bacteria can live in extreme environments can reveal useful biochemical
processes for biotechnology. Most important, finding out how bacteria cope with
contaminated sites may provide useful enzymes for cleaning up our pollution.
Bioremediation is one avenue in which biotechnology has made rapid advances. Many different humanmade compounds have
contaminated the environment around us, through everyday use, accidental
spillage, or intentional dumping. Many environmental biotechnologists are
working on “biological” means of cleaning the environment. In fact, releasing
an organism that can degrade a pollutant would provide a very easy, low-cost
way of cleaning up a polluted site.
Naturally occurring
microorganisms often have the ability to degrade humanmade pollutants. For
example, Rhodococcus has a highly
diverse repertoire of pathways to degrade pollutants, such as short- and
long-chain alkanes, aromatic molecules (both halogenated and
nitro-substituted), and heterocyclic and polycyclic aromatic compounds,
including quinolone, pyridine, thiocarbamate, s-triazine herbicides, 2-mercaptobenzothiazole (a rubber
vulcanization accelerator), benzothiophene, dibenzothiphene, MTBE (see later
discussion), and the related ethyl tert-butyl
ether (ETBE). Rhodococcus has several
features that contribute to its ability to degrade so many compounds. First, it
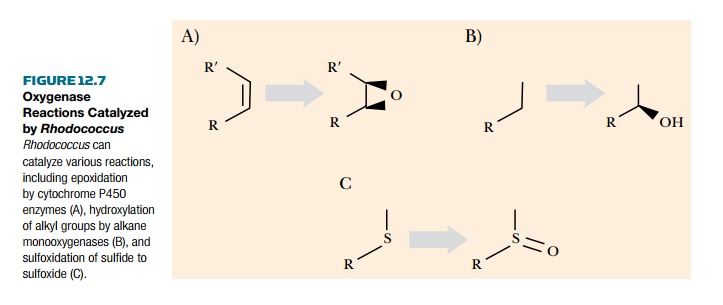
has a range of different enzymes that degrade toxic compounds, including
cytochrome P450 enzymes. These are very efficient and versatile in oxidation
pathways and catalyze a variety of reactions, including epoxidation (Fig.
12.7). Other enzymes that catalyze key degradation steps include monooxygenases
and dioxygenases, which help degrade aromatic compounds.
Furthermore, several strains of Rhodococcus can survive in solvents such as ethanol, butanol, dodecane, and toluene, which would kill many other bacteria. The oil-degrading strains actually adhere to oil droplets! Rhodococcus species are found in all types of environments, including nuclear waste sediments, tropical soil, Arctic soil, and sites in Europe, Japan, and the United States. Genetically speaking, Rhodococcus also has unique attributes that are advantageous in biodegradation. The genome of Rhodococcus sp. strain RHA1 has 9.7 Mb of DNA, including one chromosome and three large linear plasmids. The plasmids may be critical because they are important for gene transfer and recombination events. The genes for the catabolic enzymes are often found in clusters, flanked by inverted repeats, suggesting that they are acquired and passed from one strain to another by recombination. Such horizontal gene transfer can also transfer these catabolic regions to other bacteria, including Pseudomonas and Mycobacterium., Pathway Engineering, describes some plasmids that encode pollutant-degrading enzymes.
Different pollutants are
degraded in various ways. Sometimes a single naturally occurring organism can
completely degrade a pollutant. Other pollutants require more than one type of
bacterium to achieve complete degradation. Some pollutants are degraded very
slowly. Heavy metals cannot be chemically degraded, so they pose more of a
challenge than organic molecules. Most environmental biotechnologists look for
microorganisms that sequester the heavy metal in a solid phase. The conversion
of such heavy metals as uranium from an aqueous phase to a solid phase can
clean drinking water supplies. Thus, certain anaerobic microorganisms can
reduce uranium(VI) to uranium(IV) by utilizing the metal as a terminal electron
acceptor. This converts the uranium from a soluble to an insoluble form. In one
uranium-contaminated site (Old Rifle, Colorado, USA), one study actually
injected acetate into the ground water. Acetate is an electron donor that stimulates
metal-reducing bacteria to sequester uranium into the solid phase. Within 50
days, some contaminated wells had uranium concentrations lower than the
regulated level. Although these results are promising, over time the acetate
dissipated and the soluble uranium levels increased again. More research is
necessary to find a permanent means to keep uranium out of ground water.
Another recalcitrant compound that can contaminate ground water, methyl tert-butyl ether (MTBE), oxygenates gasoline so that it burns more efficiently. Many cases of MTBE contaminating groundwater have been reported. Finding a natural method to clean these sites has great applicability. The United States Environmental Protection Agency has classified MTBE as a possible human carcinogen, and drinking water must contain less than 20 to 40 µg/L. One MTBE-contaminated site in South Carolina, United States, had a large plume of MTBE-contaminated gasoline leaking from an underground storage tank at a gas station. The plume ended at a drainage ditch. The concentration of MTBE in the water was low, and in the 2-meter gap between the anaerobic and aerobic zones, the MTBE was metabolized by naturally occurring microorganisms. This led to studies that determined that MTBE could be degraded by bacteria such as Methylobium petroleophilum PM1 in areas that transition from anaerobic (anoxic) to aerobic (oxic). In fact, if anaerobic regions of the MTBE plume in South Carolina were injected with a compound that released oxygen, the concentration of MTBE decreased from 20 mg/L to 2 mg/L, suggesting that biostimulation may be a good approach to clean up contamination. Biostimulation is the release of nutrients, oxidants, or electron donors into the environment to stimulate naturally occurring microorganisms to degrade a contaminant. In other areas of MTBE contamination, the site may have to undergo bioaugmentation, that is, specific microorganisms plus their energy sources may need to be added to the site. Such microorganisms may be naturally occurring, a mixture of different organisms, or even genetically modified.
Related Topics