Botany: Classical Genetics - Intragenic gene interactions | 12th Botany : Chapter 2 : Classical Genetics
Chapter: 12th Botany : Chapter 2 : Classical Genetics
Intragenic gene interactions
Intragenic
gene interactions
Interactions take place
between the alleles of the same gene i.e., alleles at the same locus is called
intragenic or intralocus gene interaction. It includes the following:
1) Incomplete dominance
(2) Codominance
(3) Multiple alleles
(4) Pleiotropic genes
are common examples for intragenic interaction.
1. Incomplete dominance – No blending of genes
The German Botanist Carl
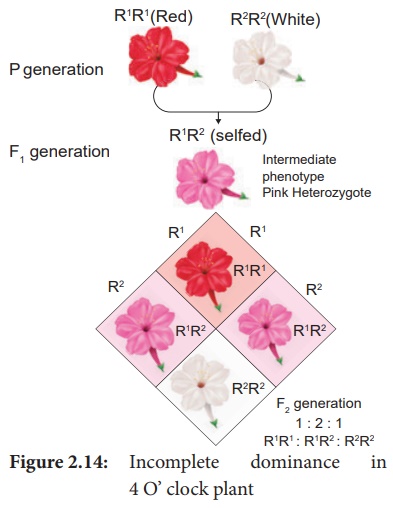
Correns’s (1905) Experiment - In 4 O’ clock plant, Mirabilis jalapa
when the pure breeding homozygous red (R1R1)
parent is crossed with homozygous white (R2R2), the
phenotype of the F1 hybrid is heterozygous pink (R1R2).
The F1 heterozygous phenotype differs from both the parental
homozygous phenotype. This cross did not exhibit the character of the dominant
parent but an intermediate colour pink. When one allele is not completely
dominant to another allele it shows incomplete dominance. Such allelic
interaction is known as incomplete dominance. F1 generation produces
intermediate phenotype pink coloured flower. When pink coloured plants of F1
generation were interbred in F2 both phenotypic and genotypic ratios
were found to be identical as 1 : 2 : 1(1 red : 2 pink : 1 white). Genotypic
ratio is 1 R1R1 : 2 R1R2 : 1 R2R2.
From this we conclude that the alleles themselves remain discrete and unaltered
proving the Mendel’s Law of Segregation. The phenotypic and genotypic ratios
are the same. There is no blending of genes. In the F2 generation R1
and R2 genes segregate and recombine to produce red, pink and white
in the ratio of 1 : 2 : 1. R1 allele codes for an enzyme responsible
for the formation of red pigment. R2 allele codes for defective enzyme.
R1 and R2 genotypes produce only enough red pigments to
make the flower pink. Two R1R1 are needed for producing
red flowers. Two R2R2 genes are needed for white flowers.
If blending had taken place, the original pure traits would not have appeared and
all F2 plants would have pink flowers. It is very clear that
Mendel’s particulate inheritance takes place in this cross which is confirmed
by the reappearance of original phenotype in F2.
How are we going to
interpret the lack of dominance and give explanation to the intermediate
heterozygote phenotype?
How will you explain
incomplete dominance at the molecular level?
Gene expression is
explained in a quantitative way. Wild-type allele which is a functional allele
when present in two copies (R1 R1) produces an functional enzyme which
synthesizes red pigments. The mutant allele which is a defective allele in two
copies (R2 R2) produces an enzyme which cannot synthesize necessary red
pigments. The white flower is due to the mutation causing complete loss of
function. The F1 intermediate phenotype heterozygote (R1R2) has one
copy of the allele R1. R1 produces 50% of the functional protein resulting in
half of the pigment of red flowered plant and so it is pink. The intermediate
phenotype pink heterogyzote with 50% of functional protein is not enough to
create the red phenotype homozygous, which makes 100% of the functional
protein.
2. Codominance (1 : 2 : 1)
This pattern occurs due
to simultaneous (joint) expression of both alleles in the heterozygote - The phenomenon in which
two alleles are both expressed in the heterozygous individual is known
as codominance. Example: Red and white flowers of Camellia, inheritance
of sickle cell haemoglobin, ABO blood group system in humanbeings. In humanbeings,
IA and IB alleles of I gene are codominant which follows Mendels law of
segregation. The codominance was demonstrated in plants with the help of
electrophoresis or chromatography for protein or flavonoid substance. Example: Gossypium
hirsutum and Gossypium sturtianum, their F1 hybrid
(amphiploid) was tested for seed proteins by electrophoresis. Both the parents
have different banding patterns for their seed proteins. In hybrids, additive
banding pattern was noticed. Their hybrid shows the presence of both the types
of proteins similar to their parents.
The heterozygote
genotype gives rise to a phenotype distinctly different from either of the
homozygous genotypes. The F1 heterozygotes produce a F2
progeny in a phenotypic and genotypic ratios of 1 : 2 : 1.
3. Lethal genes
An allele which has the
potential to cause the death of an organism is called a “Lethal Allele”. In 1907, E. Baur
reported a lethal gene in snapdragon (Antirrhinum sp.). It is an
example for recessive lethality. In snapdragon there are three kinds of plants.
1. Green plants with
chlorophyll. (CC)
2. Yellowish green
plants with carotenoids are referred to as pale green, golden or aurea plants
(Cc)
3. White plants without
any chlorophyll. (cc)
The genotype of the
homozygous green plants is CC. The genotype of the homozygous white plant is
cc.
The aurea plants have
the genotype Cc because they are heterozygous of green and white plants. When
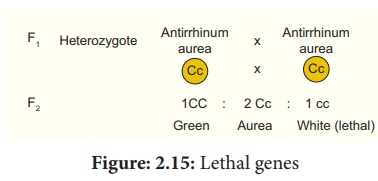
two such aurea plants are crossed the F1 progeny has identical
phenotypic and genotypic ratio of 1 : 2 : 1 (viz. 1 Green (CC) : 2 Aurea (Cc) :
1 White (cc))
Since the white plants
lack chlorophyll pigment, they will not survive. So the F2 ratio is
modified into 1 : 2. In this case the homozygous recessive genotype (cc) is
lethal.

The term “lethal” is
applied to those changes in the genome of an organism which produces effects
severe enough to cause death. Lethality is a condition in which the death of
certain genotype occurs prematurely. The fully dominant or fully recessive
lethal allele kills the carrier individual only in its homozygous condition. So
the F2 genotypic ratio will be 2 : 1 or 1 : 2 respectively.
4. Pleiotropy – A single gene affects multiple traits
In Pleiotropy, the
single gene affects multiple traits and alter the phenotype of the organism.
The Pleiotropic gene influences a number of characters simultaneously and such
genes are called pleiotropic gene. Mendel noticed pleiotropy while performing
breeding experiment with peas (Pisum sativum). Peas with purple flowers,
brown seeds and dark spot on the axils of the leaves were crossed with a
variety of peas having white flowers, light coloured seeds and no spot on the
axils of the leaves, the three traits for flower colour, seed colour and a leaf
axil spot all were inherited together as a single unit. This is due to the
pattern of inheritance where the three traits were controlled by a single gene
with dominant and recessive alleles. Example: sickle cell anemia.
Related Topics