Chapter: Plant Biochemistry: ATP is generated by photosynthesis
H -ATP synthases from bacteria, chloroplasts, and mitochondria have a common basic structure
H -ATP synthases from bacteria, chloroplasts, and mitochondria have a common basic structure
How is the energy of the proton gradient utilized to synthesize ATP?
A proton coupled ATP synthase (H –ATP+ synthase) is not unique to the chloroplast. It evolved during an early stage of evolution and occurs in its basic structure in bacteria, chloroplasts, and mitochondria. In bacteria this enzyme catalyzes not only ATP synthesis driven by a proton gradient, but also (in a reversal of this reaction) the transport of protons against the concentration gradient at the expense of ATP. This was probably the origi-nal function of the enzyme. In some bacteria an ATPase homologous to the H+ -ATP synthase functions as an ATP-dependent Na+ transporter.
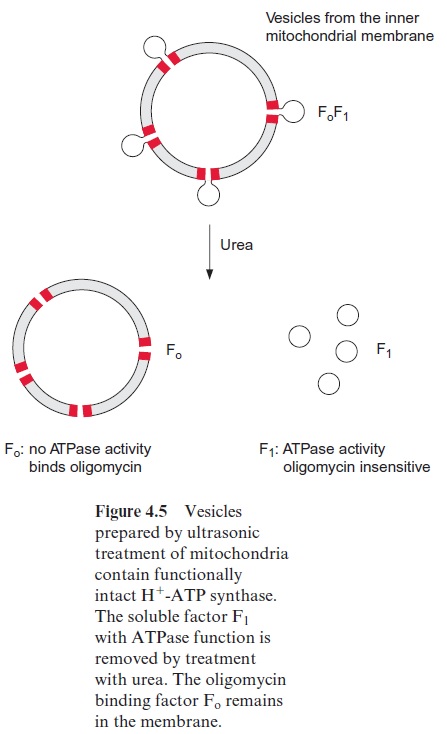
Our present knowledge about the structure and function of the H+ -ATP synthase derives from investigations of mitochondria, chloroplasts, and bacteria. By 1960 progress in electron microscopy led to the detection of small particles, which were attached by stalks to the inner membranes of mitochondria and the thylakoid membranes of chloroplasts. These particles occur only at the matrix or stromal side of the corresponding membranes. By adding urea, Ephraim Racker and coworkers (Cornell University, USA), succeeded in removing these particles from mitochondrial membranes. The isolated particles catalyzed the hydrolysis of ATP to ADP and phos-phate. Racker called them F1-ATPase. Mordechai Avron (Rehovot, Israel) showed that the corresponding particles from chloroplast membranes also have ATPase activity.
Vesicles containing F1 particles could be prepared from the inner mem-brane of mitochondria. These membrane vesicles synthesized ATP dur-ing respiration, and as in intact mitochondria , the addition of uncouplers resulted in an increased ATPase activity. The uncoupler-induced ATPase activity, as well as ATP synthesis performed by these vesicles, was found to be inhibited by the antibiotic oligomycin. Mitochondrial vesicles where the F1 particles had been removed showed no ATPase activity but were highly permeable for protons. This proton permeability was eliminated by adding oligomycin. In contrast, the ATPase activity of the removed F1 particles was not affected by oligomycin. These and other experiments showed that the H+ -ATP synthase of the mitochondria consists of two parts:
1. A soluble factor 1 (F1) that catalyzes the synthesis of ATP; and
2. A membrane-bound factor enabling the flux of protons through the membrane to which oligomycin is bound.
Racker designated this factor Fo (O, oligomycin) (Fig. 4.5). Basically the same result was found for H+ -ATP synthases of chloroplasts and bac-teria, with the exception that the H+ -ATP synthase of chloroplasts is not inhibited by oligomycin. Despite this, the membrane part of the chloro-plastic ATP synthase is also designated as Fo. The H+ -ATP synthases of chloroplasts, mitochondria, and bacteria, as well as the corresponding H+ - and Na+ -ATPases of bacteria, are collectively termed F-ATP synthasesor F-ATPases. The terms FoF1-ATP synthase and FoF1-ATPase are also used.
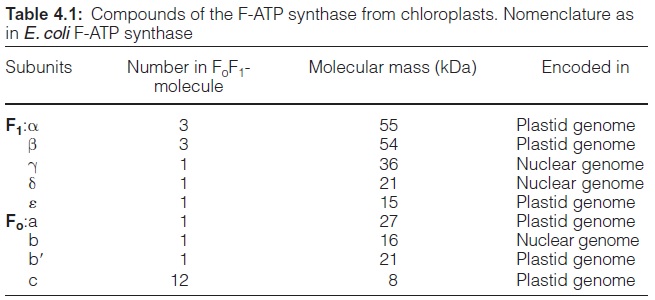
F1, after removal from the membrane, is a soluble oligomeric protein with the composition α3β3γ3δe (Table 4.1). This composition has been found in chloroplasts, bacteria, and mitochondria.
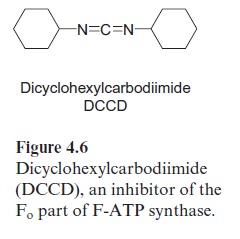
Fo is a strongly hydrophobic protein complex that can be removed from the membrane only by detergents. Dicyclohexylcarbodimide (DCCD) (Fig. 4.6) binds to the Foembedded in the membrane, and thus closes the proton channel. In chloroplasts four different subunits have been detected as the main constituents of Fo and are named a, b, b’ , and c (Table 4.1, Fig. 4.7). Subunit c, probably occurring in the chloroplastic Fo in 12–14 copies, con-tains two transmembrane helices and is the binding site for DCCD. The c subunits appear to form a cylinder, which spans the membrane. On the out-side of the cylinder spanning the membrane, the subunits a, b’, and b are arranged, where by b and b are in contact with the F1 part via subunit γ and ε form the central connection between F1 and Fo.
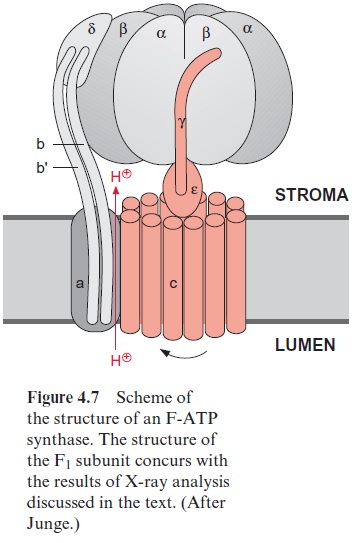
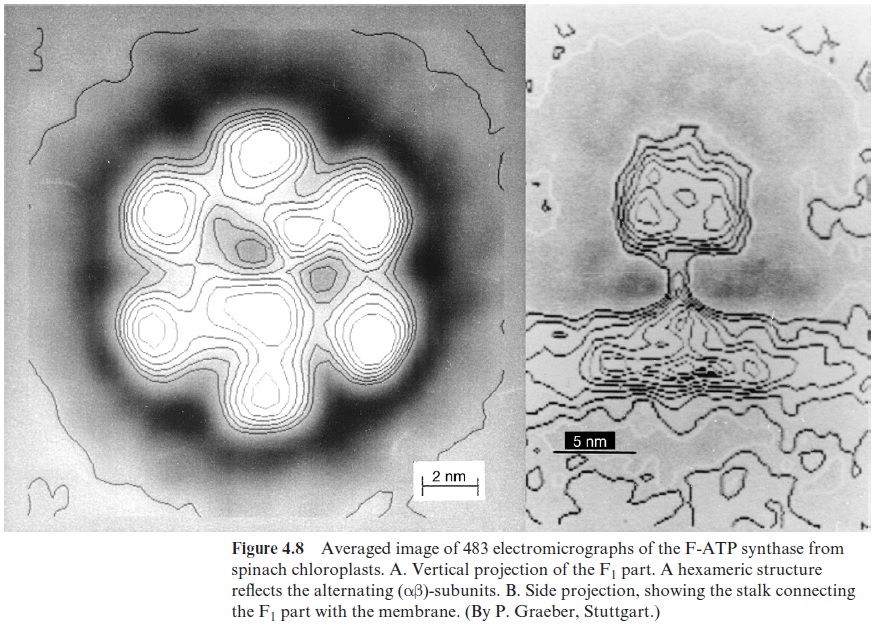
Whereas the structure of the Fo part is still somewhat hypothetical, the structure of the F1 part has been thoroughly investigated (Fig. 4.7). The F1 particles are so small that details of their structure are not visible on a single electron micrograph. However, details of the structure can be resolved if a very large number of F1 images obtained by electron microscopy are sub-jected to a computer-aided image analysis. Figure 4.8 shows a delineated image of F-ATP synthase from chloroplasts. In the side projection, the stalk connecting the F1 part with the membrane can be recognized. Using more refined picture analysis (not shown here), two stems, one thick and the other thin, were found between F1 and Fo. In the vertical projection, a hexagonal array is to be seen, corresponding to an alternating arrangement of α- and β-subunits. Investigations of the isolated F1 protein showed that an FoF1 protein has three catalytic binding sites for ADP or ATP. One of these binding sites is occupied by very tightly bound ATP, which is released only when energy is supplied from the proton gradient.
X-ray structure analysis of the F1 part of ATP synthase yields an insight into the machinery of ATP synthesis
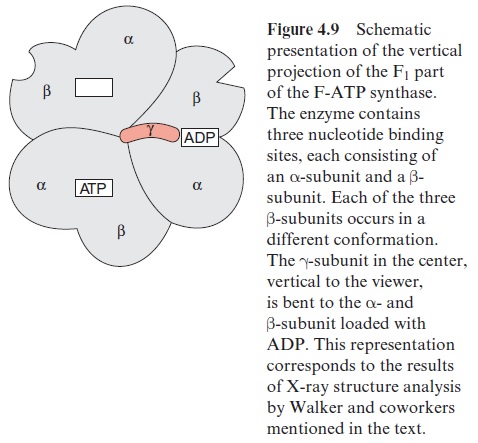
In 1994 the group of John Walker in Cambridge (England) succeeded in analyzing the three-dimensional structure of the F1 part of ATP synthase. Crystals of F1 from beef heart mitochondria were used for this analysis. Prior to crystallization, the F1 preparation was loaded with a mixture of ADP and an ATP analogue (5’ adenylyl-imidodiphosphate, AMP-PNP). This ATP analogue differs from ATP in that the last two phosphate resi-dues are connected by an N atom. It binds to the ATP binding site as ATP, but cannot be hydrolyzed by the ATPase. The structural analysis confirmed the alternating arrangement of the α- and β-subunits (Figs. 4.7 and 4.9). One α- and oneβ-subunit form a unit comprising a binding site for one adenine nucleotide. The β-subunit is primarily involved in the synthesis of ATP. In the F1 crystal investigated, one (αβ)-unit contained one ADP, the second the ATP analogue, whereas the third (αβ)-subunit was empty. These differences in nucleotide binding were accompanied by conformational dif-ferences of the three β-subunits (Fig. 4.9). The γ-subunit is arranged asym-metrically, protrudes through the center of the F1 part, and is bent to the side of the (αβ)-unit loaded with ADP (Figs. 4.7 and 4.9). This asymme-try enlightens the function of the F1 part of ATP synthase. Some general considerations about ATP synthase will be made before the function is explained in more detail.
Related Topics