Chapter: Genetics and Molecular Biology: Genes Regulating Development
Using Genetics to Begin Study of Developmental Systems
Using Genetics to Begin Study of Developmental
Systems
Although the classical experiments with insect
embryos indicated the existence of long-range effects, further progress has
required new tech-niques. Genetic approaches are one method for proceeding with
a deeper analysis of development. Mutants have the potential for indicat-ing
the complexity of a system by revealing the approximate number of genes or gene
products involved and the ways in which the system can fail. Recently,
molecular genetics has greatly streamlined the process of obtaining and
studying mutants and genes involved in the development process.
We might expect the existence of two easily
distinguishable classes of developmental mutations, maternal and embryonic.
That is, some genes and gene products necessary for spatial development likely
are expressed only in the nurse or follicle cells and are required for egg
development. Other mutations likely are embryonic and expressed only in the
developing embryo.
How can maternal lethal or embryonic lethal
mutations be isolated and studied? Certainly the technology is not as simple as
mutagenizing bacteria and providing leucine as they grow, and then identifying
those colonies that require leucine for growth. Nonetheless, the operations are
not particularly complicated either. Two problems must be solved. The first is
handling the diploid chromosomes. Most likely defects will be lethal. Since
dominant lethal mutants cannot be propagated, the mutants which can be
propagated and studied must be recessive. Recessive lethals can be generated as
long as the homologous chromo-some does not carry the mutation. Whenever the
mutation is to be detected or to be studied, both chromosomes must carry the
mutation. The second problem is preventing recombination between homologous
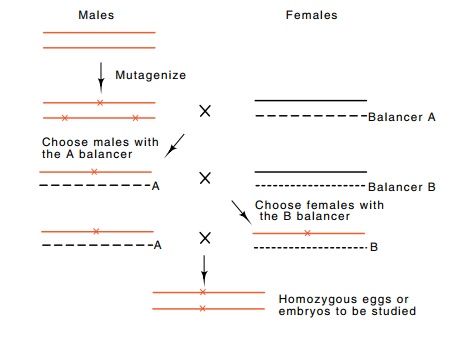
Figure
17.7 Scheme for isolating and studying
maternal and embryonic lethalmutations. A mutagenized male is mutated with a
female carrying a genetically marked balancer chromosome, indicated by B. The
genotypes of subsequent offspring can be easily identified and ultimately, a
homozygous female can be generated.
Ordinarily at meiosis, extensive pairing and
crossing over occurs between homologous chromosomes. Classical geneticists
noticed, how-ever, that if one of the chromosome pairs carries a chromosome
inver-sion, then pairing between the two chromosomes is impeded in the vicinity
of the inversion. Therefore recombination between a particular pair of
chromosomes can be greatly reduced by combining multiple inversions and
rearrangements into one of the chromosome pairs. Such a chromosome is called a
balancer chromosome.
Suppose we wish to isolate mutations in chromosome
III. The proc-ess can be started by mutagenizing a male and mating with a
female carrying a normal chromosome III and a balancer chromosome III with a
dominant, easily identified marker like wing shape or bristle pattern (Fig.
17.7). Progeny carrying the balancer will have inherited the other copy of
chromosome III from the mutagenized male. Males can be taken from this point
and used to generate females carrying the mutagenized chromosome III. Finally,
the resultant males and females carrying the mutagenized chromosome III can be
mated, and the resulting eggs or embryos lacking the dominant markers on the
balancer chromosomes can be examined for lethality or the presence of
developmental abnor-malities. In one such screen of over 5,000 mutagenized
chromosomes, about sixty maternal effect mutations were found. On average four
alleles of each of the fifteen genes were found. If the mutations were randomly
distributed among the relevant genes, these statistics imply that most of the
maternal effect genes had received at least one muta-tion. That is, that there
are on the order of 20 such genes. This is not far off. Ultimately, about 30
were found.
The general conclusions from these genetic
experiments are that a relatively small number of genes are required for
spatial development in Drosophila.
Many of the genes exert a strong effect over a sizeable portion of the embryo.
Three groups of maternal effect mutations have been found: anterior, which lack
head and thoracic structures; terminal, which lack structures at both ends; and
posterior, which lack posterior structures. Among the mutations affecting
anterior structures was the mutation known as bicoid. Embryos carrying this mutation behave similarly to embryos
with their anterior cytoplasm removed. They often lack head and thorax
structures and replace them with a mirror image of the posterior region. These
defects can be complemented by microin-jection with anterior cytoplasm from
wild-type embryos. Other muta-tions affecting head and thorax development are
the swallow and exuperantia mutations.
Additional classes of mutations that affect
embryonic pattern devel-opment independent of maternal effects have also been
found. Some of these affect segmentation in the embryo. Mutations in the gene
named fushi tarazu have too few
segments. Some mutations remove the even or the odd numbered segments, or lack
a sizeable block of segments or have segments or regions duplicated.
Developmental genes also act after segmentation.
The identities of the segments are determined by homeotic genes. For example,
the Antennapedia complex of genes
specifies development of part of thethorax, and mutants in this complex may
grow legs from the head instead of antennae. The bithorax cluster of homeotic
genes studied by Lewis specifies identity of the posterior portion of the
thorax and abdomen.
Related Topics