Chapter: Genetics and Molecular Biology: Genes Regulating Development
Expression Patterns of Developmental Genes
Expression Patterns of Developmental Genes
How can it be demonstrated that a developmental
gene actually does encode a protein and that the protein is involved in
development? Many developmental genes look like they code for proteins. The bithorax
gene complex of Drosophila helps
specify identities of segments from the second thoracic to the eighth abdominal
segment. The Ultrabithorax region
within the bithorax complex generates a number of transcripts from a very large
region containing a few exons and large introns (Fig. 17.9). The 5’ exon
contains a substantial open reading frame and looks like it codes for a
protein.
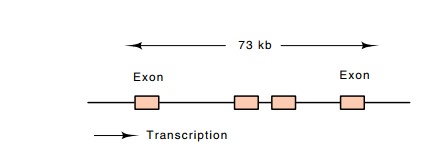
Figure
17.9 TheUbxlocus showing the distance between the exons in thetranscript.
Antibodies against the presumed ultrabithorax
protein, if they could be made, would provide proof that the gene encodes a
protein. They would also provide the basis of an assay for the presence of such
a protein so that it could be purified. The traditional approach of purify-ing
a protein and injecting it into a mouse or rabbit could hardly be used for a
protein that cannot be assayed and hasn’t been purified. Molecular genetics
techniques provided inexpensive solutions to the problem of purifying the
protein, making useful antibodies, and locating the protein within the
developing larva. A cloning vector was used from which β-galactosidase could be synthesized inE.colionly if DNA
containingan open reading frame were inserted upstream of the enzyme (Fig.
17.10). Thus Lac+ colonies were selected from cells transformed with
a plasmid into which fragments of the 5’ exon had been ligated. These would
synthesize the Ubx-β-galactosidase fusion protein, and further-more,
because of the size of the protein, it could easily be isolated from
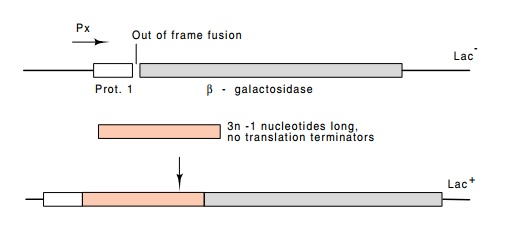
Figure
17.10 A vector for cloning stretches of
DNA containing open readingframes. Protein 1 and β-galactosidase are fused out of frame. The correct reading frame can be
restored and cells containing the vector will be Lac + if a
third segment of DNA is added which restores the correct reading frame and
which contains no translation termination codons.
SDS protein gels. β-galactosidase is one of the largest polypeptides synthesized in E. coli,
and the fusion protein likely would be even larger. Hence the fusion protein
could be separated from other proteins simply by electrophoresis. The region of
the gel containing the fusion protein was cut out, protein eluted, and injected
into rabbits or mice for direct induction of antibodies or for production of
monoclonal antibodies.
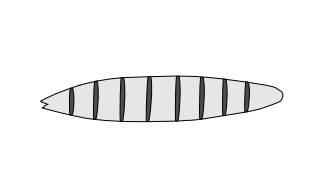
Figure 17.11 Representation ofthe pattern of expression of a ho-meotic gene in the Drosophila embryo.
The resulting antibodies against a portion of the
homeotic protein permitted assay of the developing fly for the protein’s
location and timing of synthesis. Not surprisingly, immunofluorescence shows
that the homeotic protein is found in the nuclei of cells, suggesting that it
is a DNA-binding protein. It is found in restricted parts of the embryo.
In general, in studying developmental genes, two
variables must be examined, the RNA and the protein product. We would like to
know when the mRNA is synthesized and where, and when the RNA is translated and
where the protein product goes. Questions about the RNA product can be answered
with in situ RNA-DNA hybridization
between slices of embryo and radioactive DNA. The protein products can be
located with similar in situ probing
by using antibodies directed against the protein in question. The antibodies can
be obtained as outlined above or by injecting rabbits with synthetic
polypeptides based on DNA sequences in the developmental genes.
As might be expected, the patterns of RNA and
protein synthesis from different developmental genes roughly parallel their
effects on embryos. For example, the products of genes effecting segmentation
are often expressed in stripes across the embryo (Fig. 17.11). Most of the
proteins are found in the nuclei, implying that many of the proteins regulate
gene expression directly by binding to DNA. Of course for developmental
proteins expressed after cell walls have formed, we would expect to find
developmental genes coding for secreted proteins, membrane-bound receptors, and
proteins involved in regulation cascades linking mem-brane receptors to gene
activity in the nucleus. An example of such a system is the G-protein protein
kinase pathway.
Not all proteins used early in development look
like they bind to DNA. One such example is the product of the vasa gene. This is a maternal effect
gene that is expressed only in nurse cells. It has a sequence with about 24%
amino acid identity to the translation initiation factor eIF-4A, which in
combination with 4B and 4F recognizes the cap structures on mRNA.
The bicoid mRNA and protein appears in a
particularly pleasing pattern. Bicoid mRNA is synthesized in the nurse cells
and transferred to the oocyte during oogenesis. This RNA binds just inside the
oocyte at the extreme anterior end by means of the swallow and exuperantia
gene products. The bicoid mRNA is not translated until the egg is fertilized.
This is a marvelous way for the egg to generate a gradient in the protein just
at the time it is needed. The egg can sit for a long time and diffusion cannot
destroy the gradient. These results are also consistent with the microinjection
results.
Related Topics