Chapter: Medical Microbiology: An Introduction to Infectious Diseases: Antimicrobial Resistance
Genetics of Resistance - Bacterial Resistance to Antimicrobics
Genetics of Resistance
Intrinsic Resistance
For any antimicrobic, there are bacterial species that are typically within its spectrum and those which are not. The resistance of the latter group is referred to as intrinsic or chromo-somal to reflect its inherent nature. The resistant species have features such as permeabilitybarriers, a lack of susceptibility of the cell wall, or ribosomal targets that make them inher-ently insusceptible. Some species constitutively produce low levels of inactivating enzymes, particularly the β-lactamases of Gram-negative bacteria. The chromosomal genes encoding these β-lactamases may be under repressor control and subject to induction by certain β-lactam antimicrobics. This leads to increased production of β-lactamase, which usuallyresults in resistance not only to the inducer but other β-lactams to which the organism would otherwise be susceptible. Many of the ESBLs operate in this manner.
Acquired Resistance
When an initially susceptible species develops resistance, such acquired resistance can be mutational or derived from another organism by the acquisition of new genes using one of the mechanisms of genetic exchange described. Of these, conjugation and transposition are the most important and often work in tandem.
Mutational Resistance Acquired resistance may occur when there is a crucial mutationin the target of the antimicrobic or in proteins related to access to the target (ie, perme-ability). Mutations in regulatory proteins can also lead to resistance. Mutations take place at a regular but low frequency and are expressed only if they are not associated with other effects that are disadvantageous to the bacterial cell. Mutational resistance can emerge in a single step or evolve slowly requiring multiple mutations before clinically significant resistance is achieved. Single-step mutational resistance is most likely when the antimi-crobic binds to a single site on its target. Resistance can also emerge rapidly when it is re-lated to gene regulation, such as mutational derepression of a chromosomally encoded cephalosporinase. A slow, progressive resistance evolving over years, even decades, is typical for β-lactamresistance related to altered PBPs.
Plasmids and Conjugation The transfer of plasmids by conjugation was the first dis-covered mechanism for acquisition of new resistance genes, and it continues to be the most important. Resistance genes on plasmids (R plasmids) can determine resistance to one antimicrobic or to several that act by different mechanisms. After conjugation, the re-sistance genes may remain on a recircularized plasmid or less often become integrated into the chromosome by recombination. Of course, resistance is not the only concern of plasmids. A single cell may contain more than one distinct plasmid and/or multiple copies of the same plasmid. Although most resistance mechanisms have been linked to plasmids in one species or another, plasmid distribution among the bacterial pathogens is by no means uniform. The compatibility systems that maintain plasmids from one bacteria cell generation to the next are complex. Some species of bacteria are more likely than others to contain plasmids at all. For example, Neisseria gonorrhoeae typically has multiple plasmids, whereas closely related Neisseria meningitidis rarely has any.
Plasmids are most likely to be transferred to another strain if they are conjugative, that is, if the resistance plasmid also contains the genes mediating conjugation. Another factor in the spread of plasmids is their host range. Some plasmids can be transferred only to closely related strains; others can be transferred to a broad range of species in and beyond their own genus. A conjugative plasmid with a broad host range has great potential to spread any resistance genes it carries.
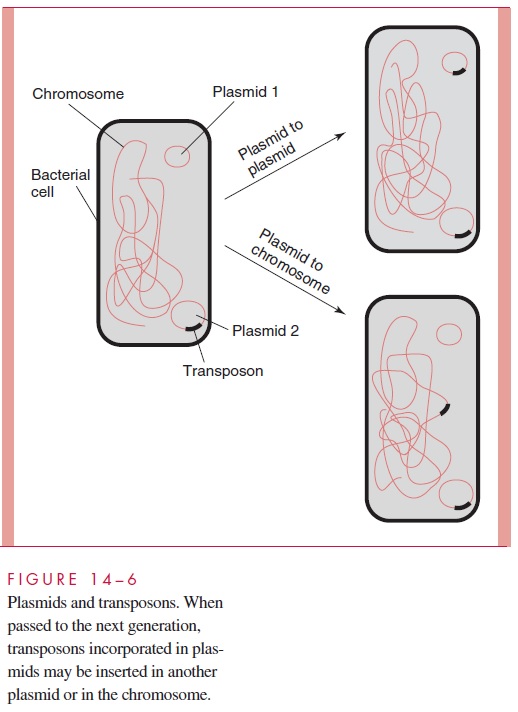
Transposons and Transposition Transposons containing resistance genes can movefrom plasmid to plasmid or between plasmid and chromosome. Many of the resistance genes carried on plasmids are transposon insertions which can be carried along with the rest of the plasmid genome to another strain by conjugation. Once there, the transposon is free to remain in the original plasmid, insert in a new plasmid, insert in the chromosome, or any combination of these (Fig 14 – 6). Theoretically, plasmids can accomplish the same events by recombination, but the nature of the transposition process is such that it is much more likely to result in the transfer of an intact gene. Transposons also have a variable host range which in general is even broader than plasmids. Together, conjugation and transposition provide extremely efficient means for spreading resistance genes.
Other Genetic Mechanisms Although the transfer of resistance genes by transductionhas been demonstrated in the laboratory, its association to clinically significant resistance has been uncommon. Transduction of imipenem resistance by wild-type bacteriophages carried by P. aeruginosa to other strains of the same bacteria is a recent example. Because of the high specificity of bacteriophages, transduction is typically limited to bacteria of the same species. Transformation is the most common way genes are manipulated in the laboratory, but detecting its occurrence in the clinical situation is particularly difficult. Plasmids are readily isolated and characterized, and transposons have flanking insertion sequences to flag their presence, but there is little to mark the uptake of naked DNA. Mol-ecular epidemiologic studies suggest that the spread of PBP mutations in Streptococcuspneumoniae is due to transformation, and there may be many more examples awaitingdiscovery.
Related Topics