Chapter: The Diversity of Fishes: Biology, Evolution, and Ecology: Systematic procedures
Taxonomic characters - Diversity of Fishes
Taxonomic characters
Whichever system of classification is employed, characters are needed to differentiate taxa and assess their interrelationships. Characters, as Stanford ichthyologist George Myers once said, are like gold – they are where you find them. Characters are variations of a homologous structure and, to be useful, they must show some variation in the taxon under study. Useful definitions of a wide variety of characters were presented by Strauss and Bond (1990).Characters can be divided, somewhat arbitrarily, into different categories.
Meristic characters originally referred to characters that correspond to body segments (myomeres), such as numbers of vertebrae and fin rays. Now, meristic is used for almostany countable structure, including numbers of scales, gill rakers, cephalic pores, and so on. These characters are useful because they are clearly definable, and usually other investigators will produce the same counts. In most cases, they are stable over a wide range of body size. Also, meristic characters are easier to treat statistically, so comparisons can be made between populations or species with aminimum of computational effort.
Morphometric characters refer to measurable structures such as fin lengths, head length, eye diameter, or ratios between such measurements. Some morphometric characters are harder to define exactly, and being continuous variables, they can be measured to different levels of precision and so are less easily repeated. Furthermore, there is the problem of allometry, whereby lengths of different body parts change at different rates with growth. Thus analysis of differences is more complex than with meristic characters. Size factors have to be compensated for through use of such techniques as regression analysis, analysis of variance (ANOVA), and analysis of covariance (ANCOVA) so that comparisons can be made between actual differences in characters and not differences due to body size. Principal components analysis (PCA) also adjusts for size, particularly if size components are removed by shear coeffi cients, as recommended by Humphries et al.(1981).
Widely used definitions of most meristic and morphometriccharacters were presented by Hubbs and Lagler (1964); some of these are illustrated in Fig. 2.2.
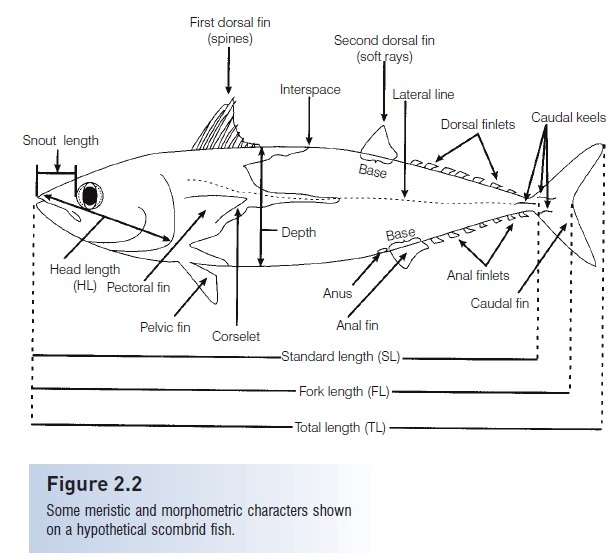
Figure 2.2
Some meristic and morphometric characters shown on a hypothetical scombrid fish.
Anatomical characters include characters of the skeleton(osteology) and characters of the soft anatomy, such as position of the viscera, divisions of muscles, and branches of blood vessels. Some investigators favor osteological characters because such characters have been thought to vary less than other characters. In some cases, this supposition has been due to the use of much smaller sample sizes than with the analysis of meristic or morphometric characters.
Other characters can include almost any fixed, describable differences among taxa. For example, color can include such characters as the presence of stripes, bars, spots, or specifi c colors. Photophores are light-producing structures that vary in number and position among different taxa. Sexually dimorphic (“two forms”) structures can be of functional value, including copulatory organs used by malesto inseminate females, like the gonopodium of a guppy(modified anal fin) or the claspers of chondrichthyans(modified pelvic fins). Cytological (including karyological),electrophoretic, serological, behavioral, and physiological characters are useful in some groups.
Molecular characters, especially nuclear DNA and mitochondrial DNA (mtDNA) have become increasingly useful at all levels of classification. All organisms contain DNA, RNA, and proteins. Closely related organisms show a high degree of similarity in molecular structures. Molecular systematics uses such data to build trees showing relationships. It is becoming easier and cheaper to sequence longer sequences of nucleotides.
Molecular data can be used to test hypotheses of relationships based on morphological data. An example are the analyses of similar morphological data sets for the Scombroidei by Collette et al. (1984) and by Johnson(1986) that produced different cladograms resulting in very different classifications. In a computer-generated cladogram(WAGNER 78; Farris 1970), Collette et al.(1984)postulated a sister-group relationship of the Wahoo (Acanthocybium)and Spanish mackerels (Scomberomorus) within the family Scombridae. In contrast, Johnson (1986) placed the Wahoo as sister to the billfishes within a greatlyexpanded Scombridae that includes billfishes as a tribe, instead of being in the separate families Xiphiidae and Istiophoridae. In part, the different authors reached different conclusions because they analyzed the data sets differently.
Another part of the differences in classification centers on the large amount of homoplasy present. No matter which classification is employed, a large number of characters must be postulated to show reversal or independent acquisition. Either more data or a different method of analysis was needed to resolve the conflict. Moleculardata, both nuclear and mitochondrial DNA (Orrell et al.2006), supports the view that the Wahoo is a scombrid and strongly refutes a close relationship between billfishes and scombroids.
Another use of molecular data is in what has been termed barcoding. This relies on differences between species in a relatively short segment of mtDNA, an approximately 655base pair region of cytochrome oxidase subunit I gene(COI) which Hebert et al. (2003) have proposed as a global bioidentification system for animals. It has been likened to the barcodes that we see on items in grocery stores. Forbarcoding to be successful, within-species DNA sequences need to be more similar to each other than to sequencesof different species. Successful barcoding will facilitate identification of fishes, linking larvae with adults, forensic identification of fishfillets and other items in commerce, and identification of stomach contents. One potential problem is that using only a mitochondrial marker may fail to discriminate between species due to introgression of some maternally inherited characters, as has apparently happened between two species of western Atlantic Spanish mackerels, Scomberomorus maculatus and S. regalis(Banford et al. 1999; Paine et al. 2007).
To test its utility in fishes, Ward et al. (2005) barcoded207 species of fishes, mostly Australian marine fishes. With no exceptions, all 207 sequenced species were discriminated. Similarly, except for one case of introgression, all17 species of western Atlantic Scombridae were successfully discriminated with COI (Paine et al. 2007). Successes like these led to ambitious plans at a 2005 workshop held at the University of Guelph in Canada to sequence all species of fishes for the Fish Barcode of Life or FISH-BOL, fostered by the Consortium for the Barcode of Life and the Census of Marine Life. This is planned to be part of a grand scheme to produce a DNA global database for all species on planet Ocean.
Related Topics