Chapter: Genetics and Molecular Biology: An Overview of Cell Structure and Function
Diffusion within the Small Volume of a Cell
Diffusion within the Small Volume of a Cell
Within
several minutes of adding a specific inducer to bacteria or eukaryotic cells,
newly synthesized active enzymes can be detected. These are the result of the
synthesis of the appropriate messenger RNA, its translation into protein, and
the folding of the protein to an active conformation. Quite obviously,
processes are happening very rapidly within a cell for this entire sequence to
be completed in several minutes. We will see that our image of synthetic
processes in the cellular interior should be that of an assembly line running
hundreds of times faster than normal, and our image for the random motion of
molecules from one point to another can be that of a washing machine similarly
running very rapidly.
The
random motion of molecules within cells can be estimated from basic physical
chemical principles. We will develop such an analysis since similar reasoning
often arises in the design or analysis of experi -
ments in molecular biology. The mean squared distance Bar R2 that a molecule with diffusion
constant D will diffuse in time t is Bar R2= 6Dt (Fig. 1.10). The diffusion constants of many molecules have been
measured and are available in tables. For our purposes, we can estimate a value for a diffusion constant.
The diffusion constant is D=KTŌüäf , where K
is the Boltzmann constant, 1.38 ├Ś 10-16 ergs/degree, T is temperature in degrees Kelvin, and f is the frictional force. For spherical bodies, f =6ŽĆ╬Ęr, where r is the radius in centimeters and╬Ęis the viscosity ofthe medium in
units of poise.
The
viscosity of water is 10-2 poise. Although the macroviscosity of the
cellŌĆÖs interior could be much greater, as suggested by the extremely high
viscosity of gently lysed cells, the viscosity of the cellŌĆÖs interior with
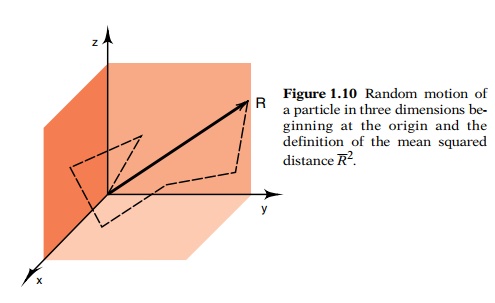
Figure
1.10 Random motion ofa particle in
three dimensions be-ginning at the origin and the
respect
to motion of molecules the size of proteins or smaller is more likely to be
similar to that of water. This is reasonable, for small molecules can go around
obstacles such as long strands of DNA, but large molecules would have to
displace a huge tangle of DNA strands. A demonstration of this effect is the
finding that small molecules such as amino acids readily diffuse through the
agar used for growing bacterial colonies, but objects as large as viruses are
immobile in the agar, yet diffuse normally in solution.
Since D=KTŌüä6ŽĆ╬Ęr, then D = 4.4 ├Ś 10-7 for a large
spherical protein of radius 50 ├ģ diffusing in water, and the diffusion constant
for such a protein__ within a
cell is not greatly different. Therefore R2=6├Ś4.4├Ś10ŌłÆ7t, and the
average time required for such proteinmolecules to diffuse the length of a 1 ┬Ą bacterial cell is 1/250 second
and to diffuse the length of a 20 ┬Ą eukaryotic cell is about 2 seconds. Analogous
reasoning with respect to rotation shows that a protein rotates about 1/8
radian (about 7┬░) in the time it diffuses a distance equal to its radius.
Related Topics