Chapter: Modern Pharmacology with Clinical Applications: β-Lactam Antibiotics
β-Lactam Antibiotics: Mechanisms of Resistance
MECHANISMS OF
RESISTANCE
A number of microorganisms
have evolved mecha-nisms to overcome the inhibitory actions of the β-lactam
antibiotics. There are four major mechanisms of resistance: inactivation of the
β-lactam ring, alteration of PBPs, reduction of antibiotic access to PBPs, and
elaboration of antibiotic efflux mechanisms. Bacterial resistance may arise
from one or more than one of these mechanisms.
The most important mechanism
of resistance is hy-drolysis of the β-lactam ring by β-lactamases
(penicilli-nases and cephalosporinases). Many bacteria (Staphylo-coccus aureus, Moraxella [Branhamella] catarrhalis,
Neisseria gonorrhoeae, Enterobacteriaceae, Haemophilus influenzae, and
Bacteroides spp.) possess β-lactamases
that hydrolyze penicillins and cephalosporins. The β-lactamases evolved
from PBPs and acquired the capacity to bind β-lactam antibiotics, form an acyl
enzyme mole-cule, then deacylate and hydrolyze the β-lactam ring. Some bacteria
have chromosomal (inducible) genes for β-lactamases. Other bacteria acquire β-lactamase
genes via plasmids or transposons. Transfer of β-lactamase genes between
bacterial species has contributed to the proliferation of resistant organisms
resulting in the ap-pearance of clinically important adverse consequences.
Efforts to overcome the
actions of the β-lactamases have led to the development of such β-lactamase
in-hibitors as clavulanic acid, sulbactam, and tazobactam. They are called
suicide inhibitors because they perma-nently bind when they inactivate β-lactamases.
Among the β-lactamase inhibitors, only clavulanic acid is avail-able for oral use.
Chemical inhibition of β-lactamases, however, is not a permanent solution to
antibiotic resistance, since some β-lactamases are resistant to clavulanic
acid, tazobactam, or sulbactam. Enzymes re-sistant to clavulanic acid include
the cephalosporinases produced by Citrobacter
spp., Enterobacter spp., and Pseudomonas aeruginosa.
An additional mechanism of
antibiotic resistance in-volves an alteration of PBPs. Resistant bacteria,
usually gram-positive organisms, produce PBPs with low affin-ity for β-lactam
antibiotics. The development of muta-tions of bacterial PBPs is involved in the
mechanism for β-lactam resistance in Streptococcus
pneumoniae, Enterococcus faecium, and
methicillin-resistant S. au-reus (MRSA).
Some gram-negative bacteria
employ a third mech-anism of resistance, namely, one that reduces antibiotic
access to PBPs. Gram-positive organisms have an ex-posed peptidoglycan layer
easily accessible to β-lactam antibiotics (Fig. 45.2). In contrast, gram-negative
organ-isms have an outer membrane surrounding the peptido-glycan layer.
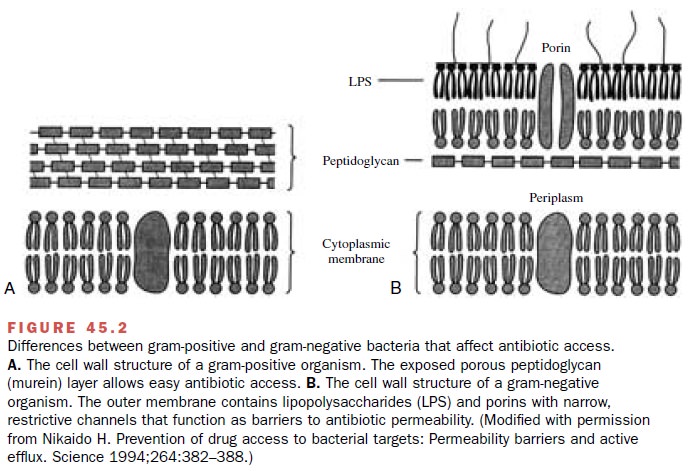
The gram-negative outer membrane hin-ders ingress of large molecules and helps bacteria resist the actions of antibiotics. In susceptible gram-negative bacteria, protein channels
(porins) allow β-lactam an-tibiotics to traverse the outer membrane and
interact with PBPs in the periplasmic space. In resistant bacteria like P. aeruginosa, porin mutants impede the β-lactam
transfer across the outer membrane.
Finally, some gram-negative
organisms demonstrate a fourth mechanism of resistance. For example, strains of
P. aeruginosa produce xenobiotic
efflux pumps to eject antibiotics. Drug efflux mechanisms are associated with
multiple drug resistance, including resistance to β-lactam antibiotics.
Widespread use of β-lactam
antibiotics exerts selec-tive pressure on bacteria and permits the
proliferation of resistant organisms. A comparison of current antibi-ograms
with those from previous decades shows an alarming increase in bacterial
resistance to β-lactam an-tibiotics.
Related Topics