Chapter: Pharmaceutical Biotechnology: Fundamentals and Applications : Genomics, Other “Omics” Technologies, Personalized Medicine, and Additional Biotechnology Related Techniques
“Omics” Integrating Technology: Systems Biology
“Omics” Integrating Technology: Systems Biology
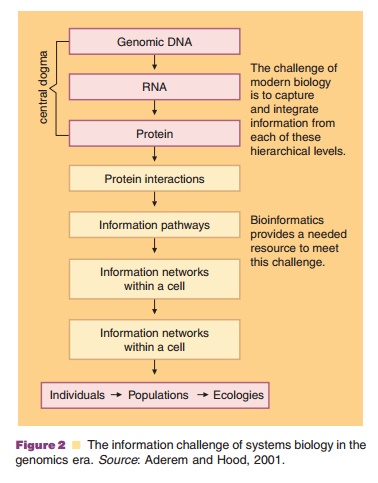
The massive scientific effort embodied in the HGP and the development of bioinformatics technologies have catalyzed fundamental changes in the practice of modern biology (Aderem and Hood, 2001). Biology has become an information science defining all the elements in a complex biological system and placing them in a database for comparative interpretation. As seen in Figure 2, the hierarchy of information collec-tion goes well beyond the biodata contained in the genetic code that is transcribed and translated. It involves a complex interactive system. Systems biol-ogy is often described as a non-competitive technology by the pharmaceutical industry (a foundational technology that must be developed to better succeed at the competitive technology of drug discovery and development). It is the study of the interactions between the components of a biological system, and how these interactions give rise to the function and behavior of that system. The biological system may involve enzymes and metabolites in a metabolic pathway or other interacting biological molecules affecting a biological process. Characterized by a cycle of theory, computational modeling and experiment to quantitatively describe cells or cell processes, systems biology is a data intensive endeavor (Klipp et al., 2005; Rothberg et al., 2005). Since the objective is a model of all the interactions in a system, the experimental techniques that most suit systems biology are those that are system-wide and attempt to be as complete as possible. High-throughput omics technologies such as proteomics, pharmacogenomics, transcriptomics, me-tabolomics, and toxicogenomics are used to collect quantitative data for the construction and validation of systems models.
Other “Omics” Technologies
The burgeoning fields of genomics, proteomics, etc. have spawned an ever increasing number of sub-disciplines and related specialty areas of study. The complexity in terminology for “omic” technologies has amplified even further with the introduction of terms such as interactomics, proteogenomics, nutri-genomics, etc. Interactomics is the data intensive broad system study of the interactome, which is the interaction among proteins and other molecules within a cell. Proteogenomics has been used as a broadly encompassing term to describe the merging of genomics, proteomics, small molecules, and infor-matics. Nutrigenomics is a new field evaluating the response of living organisms to the presence or absence of important nutrients. Lipidomics is ob-viously the large-scale study of all non-water-soluble lipid metabolites in an organism. The “omics” explo-sion will continue.
Related Topics